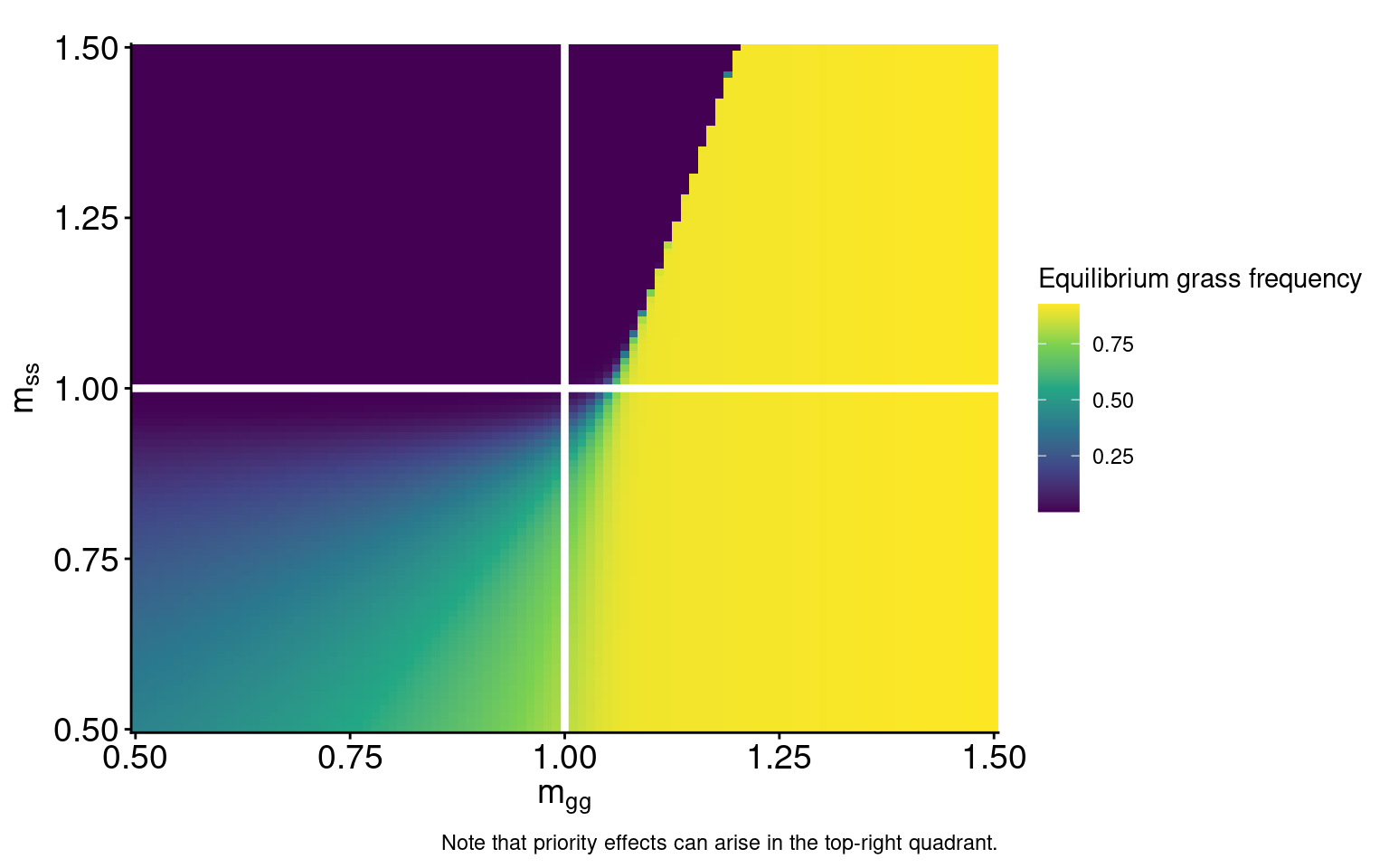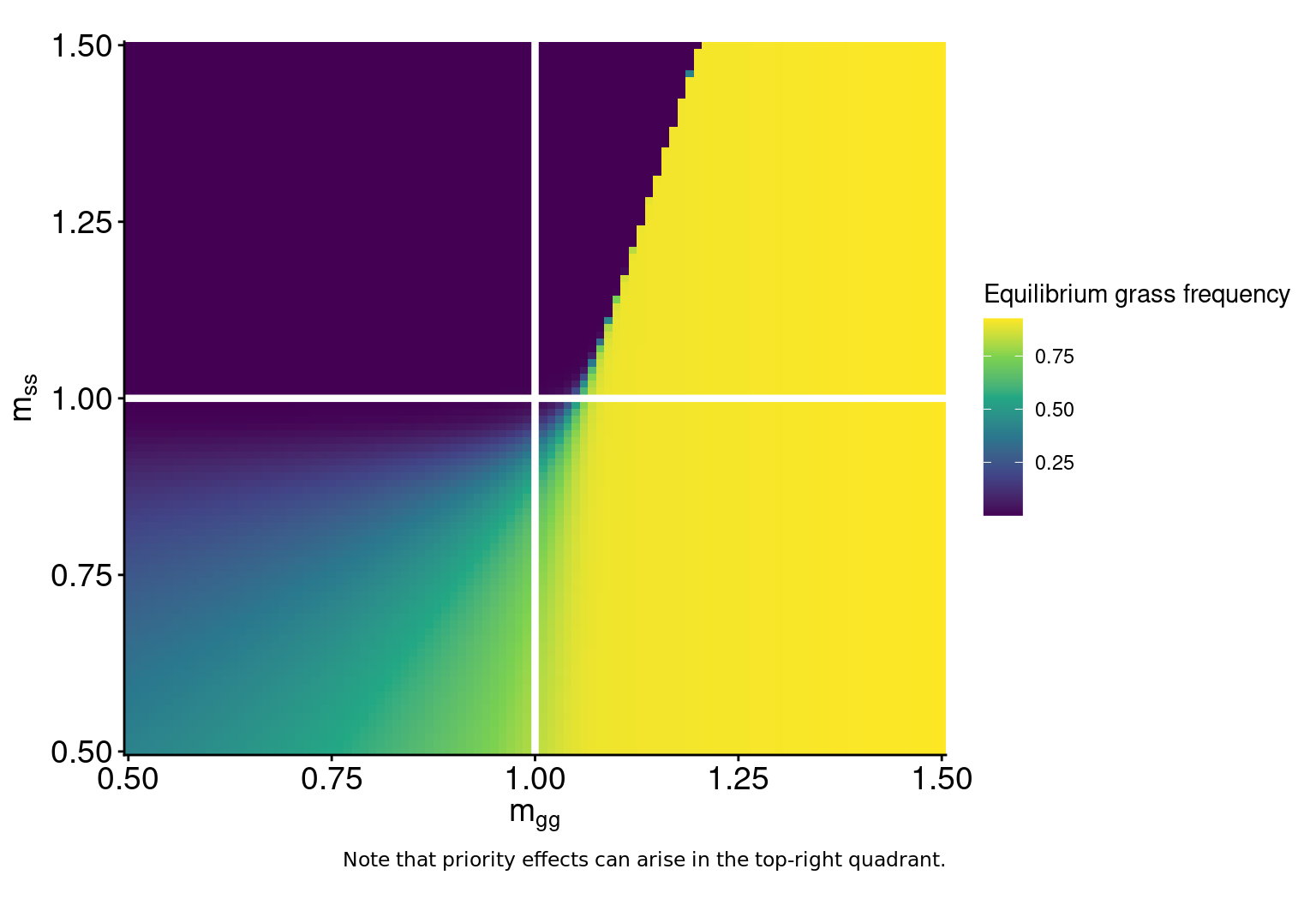Belowground regulation of
aboveground dynamics
Gaurav Kandlikar
Louisiana State University
web: https://gklab.org/
slides: https://talks.gklab.org/umn-26
contact: gkandlikar@lsu.edu
Sal (Shorea) dominated forests in northern India, Photo by Dhritiman Mukherjee
Intercropping of soybean and wheat; image from U. Iowa
“Christmas tree scientists work to manage grinchy fungal foes”; image from Washington State Univ
As an ecologist, my research is motivated by two overarching goals:
Explain patterns of diversity in nature
Predict responses to perturbations
Approach


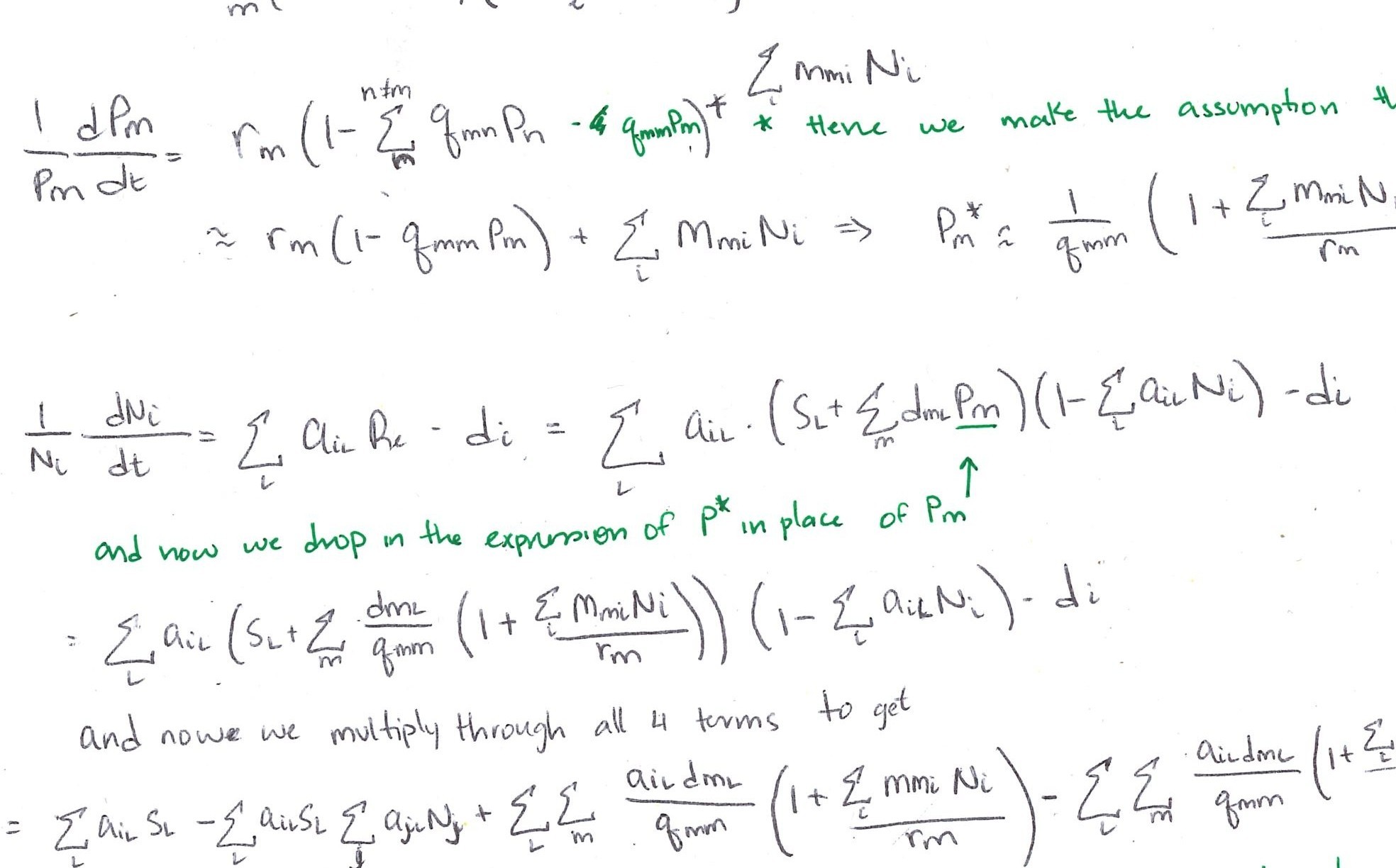
Today’s talk
Advancing ecological theory and experiments to quantify microbial effects on plant coexistence
Microbial effects on plant communities in variable environments: Ongoing work and new directions
Reflections on broadening representation and equity
Today’s talk
- Advancing ecological theory and experiments to quantify microbial effects on plant coexistence
Mathematical underpinning of plant–soil feedback
See also Bever, Westover, and Antonovics (1997) and Kandlikar (2024)
Plant dynamics
\[\frac{dN_1}{dt} = w_1N_1 ~~~~ \text{and} ~~~ \frac{dN_2}{dt} = w_2N_2\]
\[w_i = m_{iA}f_A + m_{iB}(1-f_A)\]
where \(f_i\) is the relative frequency of microbial community \(i\) in the soil (e.g. \(f_A = \frac{N_A}{N_A+N_B}\))
Interpretation: Plants have exponential population growth at a rate determined by the composition of the soil community.
Microbe dynamics
\[\frac{dN_A}{dt} = N_A\frac{N_1}{N_1 + N_2}~~~\text{and}~~~\frac{dN_B}{dt} = \nu N_B\frac{N_2}{N_1 + N_2}\]
Interpretation: Soil communities grow at a rate determined by the frequency of each plant (and the relative strength of each plant’s conditioning effect, \(\nu\)).
R code to simulate model dynamics
Code
psf_model <- function(time, init, params) {
with (as.list(c(time, init, params)), {
# description of parameters (see Bever et al. 1997)
# N1 and N2: abundance of the plant species 1 and 2
# p1: frequency of plant species 1; p2 = 1-pA
# m1A, m1B, m2A, m2B: conspecific and heterospecific effects of microbial community A or B on the growth of plant 1 or 2
# pA: frequency of the soil microbial community A
# v: influence of plant species 2 on the microbial community relative to that of plant 1
# Differential equations
dN1 <- (m1A*pA + m1B*(1-pA))*N1
dN2 <- (m2A*pA + m2B*(1-pA))*N2
dp1 <- p1*(1-p1)*((m1A-m2A)*pA + (m1B-m2B)*(1-pA))
dpA <- pA*(1-pA)*(p1-v*(1-p1))
# Return dN1 and dN2
return(list(c(dN1, dN2, dp1, dpA)))
})
}\[ \overbrace{(m_{1B} + m_{2A})}^{\text{interspecific effects}}- \overbrace{(m_{1A} + m_{2B})}^{\text{intraspecific effects}} > 0 \]
\[ \underbrace{\frac{1}{2}\big[\overbrace{(m_{1B} + m_{2A})}^{\text{interspecific effects}}- \overbrace{(m_{1A} + m_{2B})}^{\text{intraspecific effects}}\big]}_{\text{Stablization}} > 0 \]
Lasting influence of the Bever model
Code
# Dataset downloaded from Figshare:
# curl::curl_download("https://figshare.com/ndownloader/files/14874749", destfile = "../crawford-supplement.xlsx")
read_excel("../data/crawford-supplement.xlsx", sheet = "Data") |>
mutate(stab = -0.5*rrIs) |> # show in terms of stabilization instead of I_s
filter(stab > -3) |> # Remove the pair with Stab approx. 6 -- according to McCarthy Neuman (original author), this is likely due to bad seed survival
ggplot(aes(x = stab)) +
geom_histogram(color = "black") +
geom_histogram(fill = "#56A0D3") +
geom_rug(color = "grey25") +
geom_vline(xintercept = 0) +
annotate("text", x = Inf, y = Inf, label = "N = 1037 pairwise comparisons\n
57% experience positive stabilization",
vjust=1, hjust = 1, size = 6) +
xlab("Stabilization") + ylab("Count")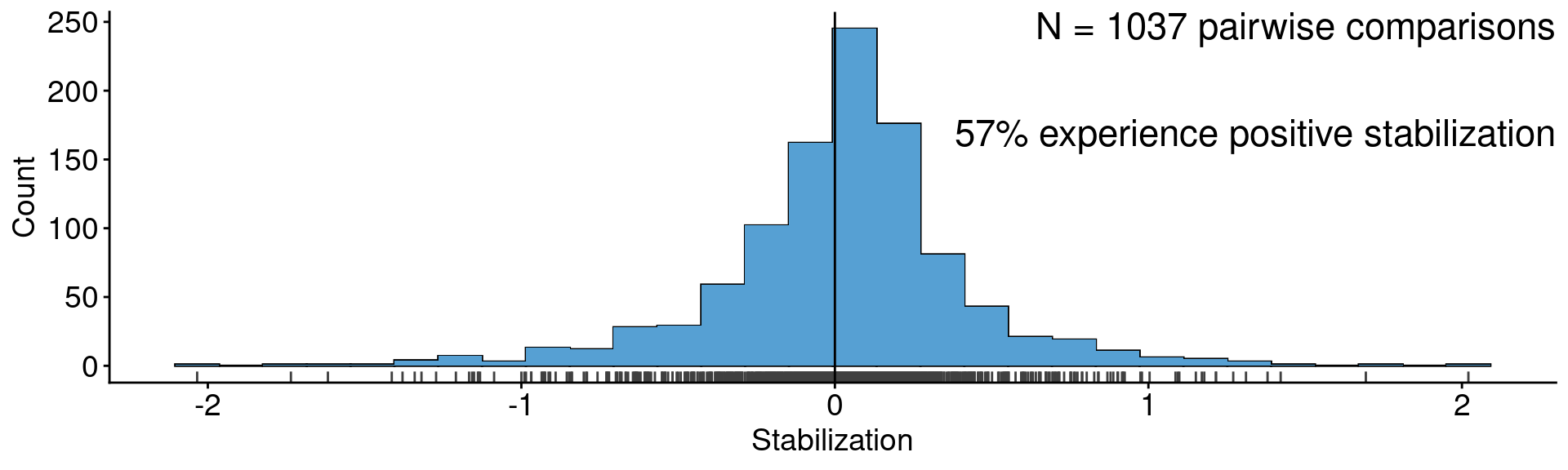
Data from Crawford et al. (2019)
Microbial feedbacks drive community patterns?
Code
mangan <-
tribble(~IS, ~abun, ~where,
-0.15504664970313828, 0.7732558139534884, "BCI",
-0.0887192536047498, 1.005813953488372, "BCI",
-0.08583545377438509, 1.4825581395348837, "BCI",
-0.09669211195928756, 1.7267441860465116, "BCI",
-0.04512298558100089, 2.1104651162790695, "BCI",
-0.006276505513146763, 2.4186046511627906, "BCI",
-0.18231106613816006, 0.3003106037506997, "Gigante",
-0.1717428991693779, 1.0202629577384568, "Gigante",
-0.13691306110550813, 1.083509520825865, "Gigante",
-0.1250429718781308, 1.425445047930817, "Gigante",
-0.09753373250550346, 1.9673983333947622, "Gigante") |>
mutate(stab = IS*-0.5)
mangan_rug_blank <-
mangan |>
ggplot(aes(x = stab,y = abun, color = where)) +
# geom_rug(sides = 'b', linewidth = 2) +
# geom_point(size = 4, stroke = 2, shape = 21) +
# geom_smooth(method = "lm", se = F) +
geom_point(color = 'transparent') +
scale_color_manual(values = c("#117733", "#cc6677")) +
geom_vline(xintercept = 0, linetype = 'dashed', color = 'grey', linewidth = 2) +
xlab("Strength of feedback (species' average stablization)") +
ylab("Log abundance of trees") +
theme(legend.position = 'none') +
scale_x_continuous(limits = c(-0.1, 0.1))+
scale_y_continuous(limits = c(-0.1,2.5))
mangan_rug_blank
Data from Mangan et al. (2010)
Microbial feedbacks drive community patterns?
Code
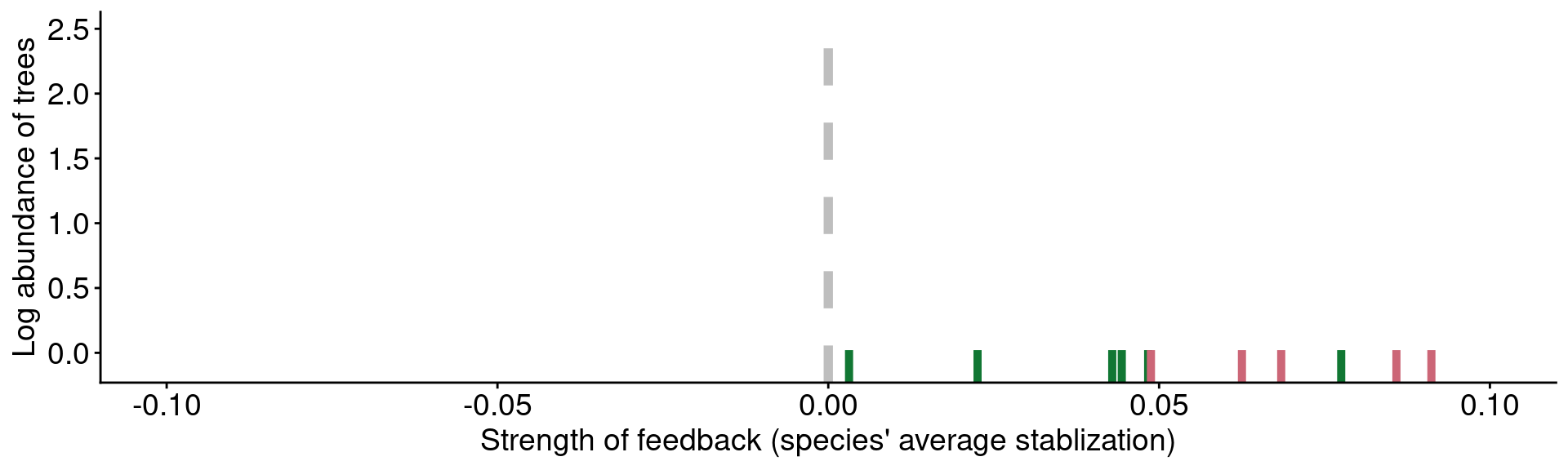
Data from Mangan et al. (2010)
Microbial feedbacks drive community patterns?
Code
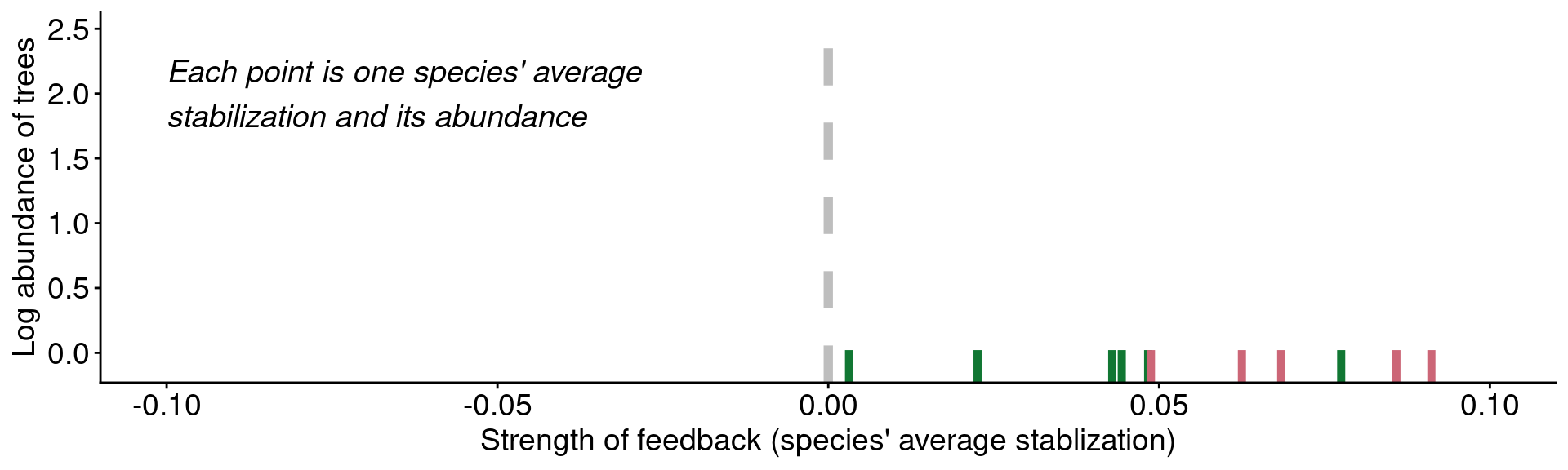
Data from Mangan et al. (2010)
Microbial feedbacks drive community patterns?
Code
Data from Mangan et al. (2010)
Microbial feedbacks drive community patterns?
Code
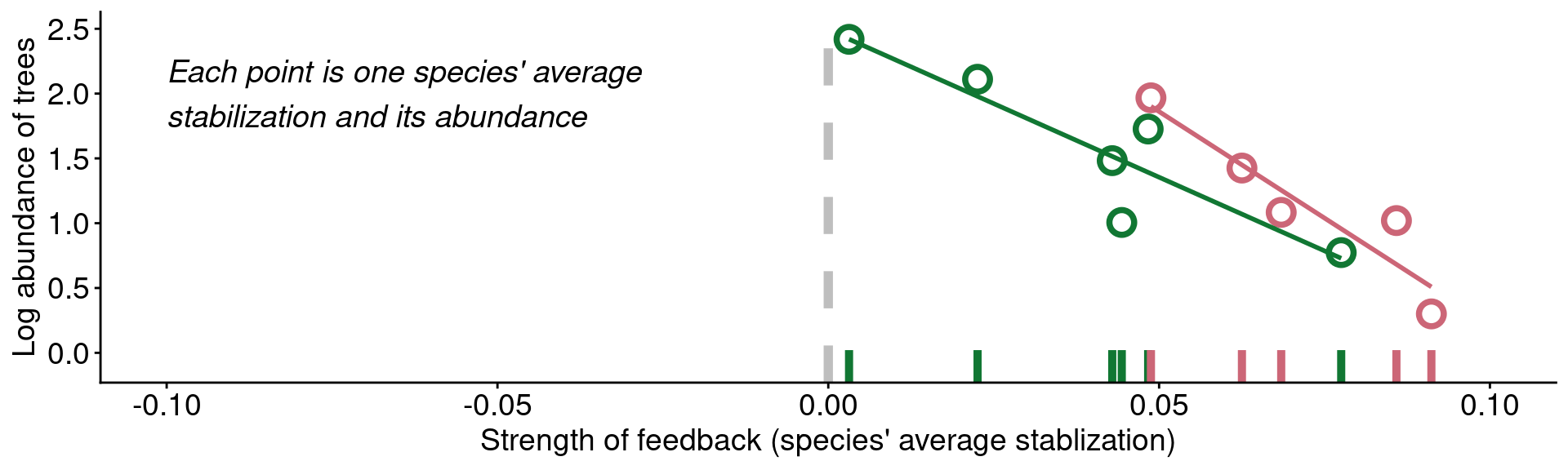
mangan_rug +
geom_point(size = 4, stroke = 2, shape = 21) +
geom_smooth(method = "lm", se = F) +
annotate('text', x = -0.1, y = 2, fontface = "italic", hjust = 0, vjust = 0.5, size = 5,
label = '"[...] trees are abundant because they are less\nsusceptible to the detrimental effects of\ntheir associated soil communities than are\nrarer tree species"')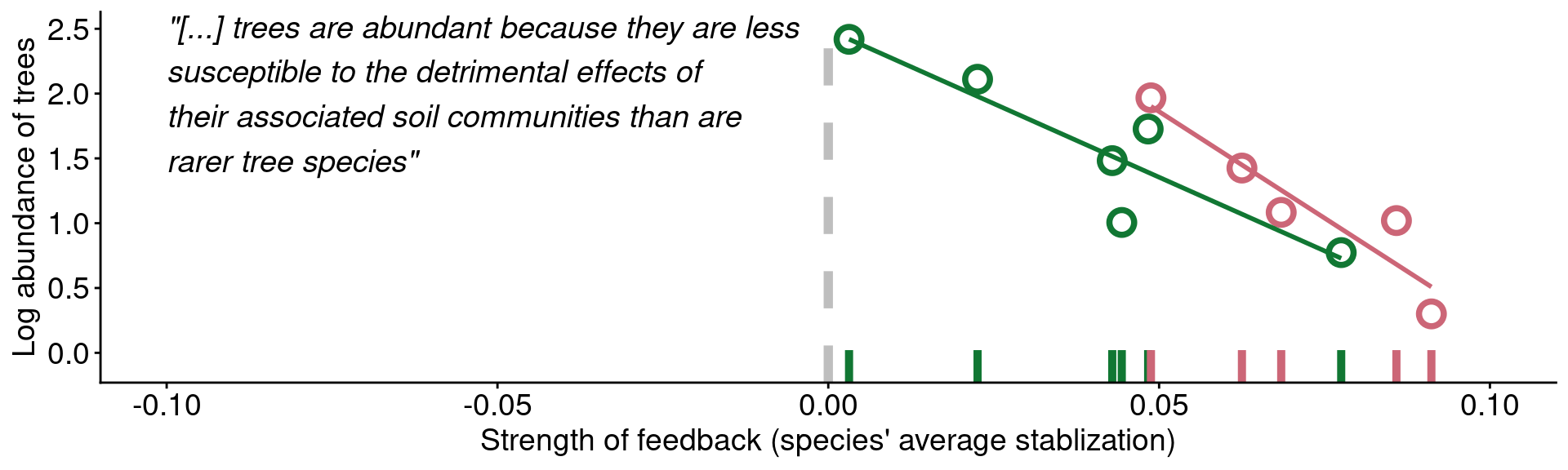
Data from Mangan et al. (2010)
A puzzle: same stabilization, different model dynamics
Code
# Define a function used for making the PSF framework
# schematic, given a set of parameter values
make_params_plot <- function(params, scale = 1.5) {
color_func <- function(x) {
ifelse(x < 0, "#EE6677", "#4477AA")
}
df <- data.frame(x = c(0,0,1,1),
y = c(0,1,0,1),
type = c("M", "P", "M", "P"))
params_plot <-
ggplot(df) +
annotate("text", x = 0, y = 1.1, size = 3.25, label = "Plant 1", color = "black", fill = "#9970ab", fontface = "bold") +
annotate("text", x = 1, y = 1.1, size = 3.25, label = "Plant 2", color = "black", fill = "#5aae61", fontface = "bold") +
annotate("text", x = 0, y = -0.15, size = 3.25, label = "Soil\nmicrobes A", label.size = 0, fill = "transparent") +
annotate("text", x = 1, y = -0.15, size = 3.25, label = "Soil\nmicrobes B", label.size = 0, fill = "transparent") +
annotate("text", x = 0.45, y = -0.4, size = 3.5, fill = "transparent",
label.size = 0.05,
label = (paste0("Stabilization = ",
0.5*(params["m1B"] + params["m2A"] - params["m1A"] - params["m2B"])))) +
geom_segment(aes(x = 0, xend = 0, y = 0.1, yend = 0.9),
arrow = arrow(length = unit(0.03, "npc")),
linewidth = abs(params["m1A"])*scale,
color = alpha(color_func(params["m1A"]), 1)) +
geom_segment(aes(x = 0.05, xend = 0.95, y = 0.1, yend = 0.9),
arrow = arrow(length = unit(0.03, "npc")),
linewidth = abs(params["m2A"])*scale,
color = alpha(color_func(params["m1B"]),1)) +
geom_segment(aes(x = 0.95, xend = 0.05, y = 0.1, yend = 0.9),
arrow = arrow(length = unit(0.03, "npc")),
linewidth = abs(params["m1B"])*scale,
color = alpha(color_func(params["m2A"]), 1)) +
geom_segment(aes(x = 1, xend = 1, y = 0.1, yend = 0.9),
arrow = arrow(length = unit(0.03, "npc"),),
linewidth = abs(params["m2B"])*scale,
color = alpha(color_func(params["m2B"]), 1)) +
# Plant cultivation of microbes
geom_segment(aes(x = -0.25, xend = -0.25, y = 0.9, yend = 0.1), linewidth = 0.15, linetype = 1,
arrow = arrow(length = unit(0.03, "npc"))) +
geom_segment(aes(x = 1.25, xend = 1.25, y = 0.9, yend = 0.1), linewidth = 0.15, linetype = 1,
arrow = arrow(length = unit(0.03, "npc"))) +
annotate("text", x = 0, y = 0.5,
label = (paste0("m1A = ", params["m1A"])),
angle = 90, vjust = -0.25, size = 3) +
annotate("text", x = 1.15, y = 0.5,
label = (paste0("m2B = ", params["m2B"])),
angle = -90, vjust = 1.5, size = 3) +
annotate("text", x = 0.75, y = 0.75,
label = (paste0("m2A = ", params["m2A"])),
angle = 45, vjust = -0.25, size = 3) +
annotate("text", x = 0.25, y = 0.75,
label = (paste0("m1B = ", params["m1B"])),
angle = -45, vjust = -0.25, size = 3) +
xlim(c(-0.4, 1.4)) +
coord_cartesian(ylim = c(-0.25, 1.15), clip = "off") +
theme_void() +
theme(legend.position = "none",
plot.caption = element_text(hjust = 0.5, size = 10))
return(params_plot)
}
# Define a function for simulating the dynamics of the
# PSF model with deSolve
params_coex <- c(m10 = 0.16, m20 = 0.16,
m1A = 0.11, m1B = 0.26,
m2A = 0.27, m2B = 0.13, v = 1)
time <- seq(0,50,0.1)
init_pA_05 <- c(N1 = 3, N2 = 7, p1 = 0.3, pA = 0.3)
out_pA_05 <- ode(y = init_pA_05, times = time, func = psf_model, parms = params_coex) |> data.frame()
params_to_plot_coex <- c(m1A = unname(params_coex["m1A"]-params_coex["m10"]),
m1B = unname(params_coex["m1B"]-params_coex["m10"]),
m2A = unname(params_coex["m2A"]-params_coex["m20"]),
m2B = unname(params_coex["m2B"]-params_coex["m20"]), v=1)
param_plot_coex <-
make_params_plot(params_to_plot_coex, scale = 6) +
annotate("text", x = 1.375, y = 0.5,
label = paste0("v = ", params_coex["v"]),
angle = -90, vjust = 1.5, size = 3)
panel_abund_coex <-
out_pA_05 |>
as_tibble() |>
select(time, N1, N2) |>
pivot_longer(N1:N2) |>
ggplot(aes(x = time, y = value, color = name)) +
geom_line(linewidth = 0.9) +
scale_color_manual(values = c("#9970ab", "#5aae61"),
name = "Plant species", label = c("Plant 1", "Plant 2"), guide = "none") +
scale_y_continuous(breaks = c(2e4, 4e4, 6e4, 8e4), labels = scales::scientific) +
ylab("Abundance") +
theme(axis.title = element_text(size = 10))
# Panel D: plot for frequencies of both plants when growing
# in dynamic soils
panel_freq_coex <-
out_pA_05 |>
as_tibble() |>
mutate(p2 = 1-p1) |>
select(time, p1, p2) |>
pivot_longer(p1:p2) |>
ggplot(aes(x = time, y = value, color = name)) +
geom_line(linewidth = 1.2) +
scale_color_manual(values = c("#9970ab", "#5aae61"),
name = "Plant species", label = c("Plant 1", "Plant 2")) +
scale_y_continuous(limits = c(0,1), breaks = c(0, 0.5, 1)) +
ylab("Frequency") +
theme(axis.title = element_text(size = 10))
param_plot_coex + {panel_abund_coex+panel_freq_coex &
theme(axis.text = element_text(size = 8), legend.position = 'none')} +
plot_layout(widths = c(1/3, 2/3))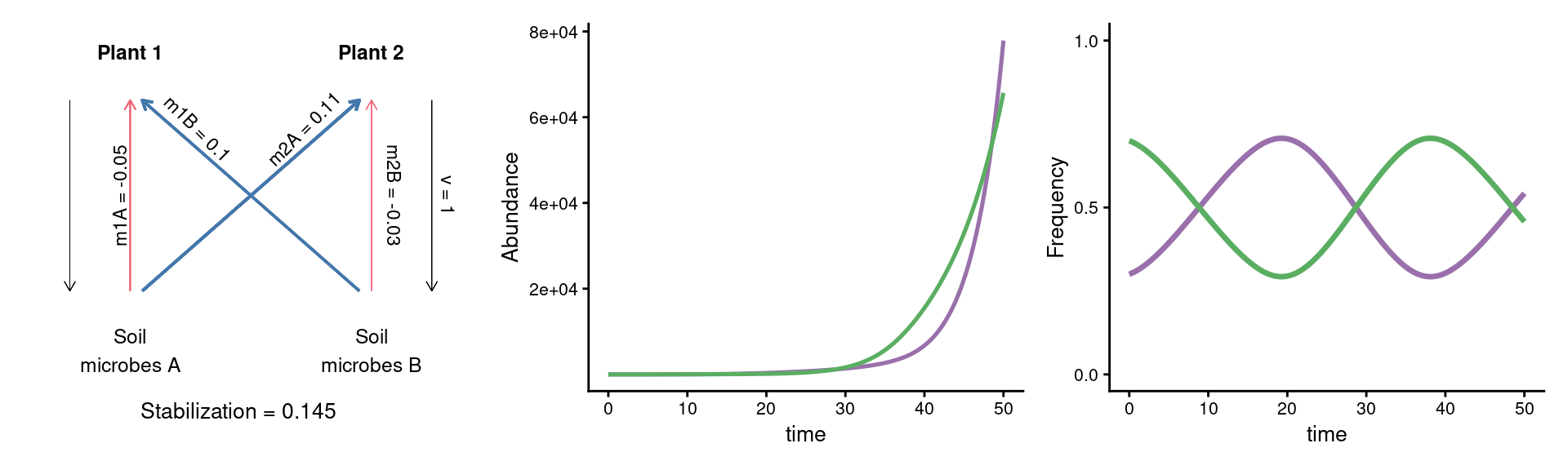
Code
# Define a function for simulating the dynamics of the
# PSF model with deSolve
params <- c(m10 = 0.16, m20 = 0.16,
m1A = 0.02, m2A = 0.33,
m1B = 0.18, m2B = 0.20, v = 1)
time <- seq(0,200,0.1)
init_pA_05 <- c(N1 = 3, N2 = 7, p1 = 0.3, pA = 0.3)
out_pA_05 <- ode(y = init_pA_05, times = time, func = psf_model, parms = params) |> data.frame()
params_to_plot <- c(m1A = unname(params["m1A"]-params["m10"]),
m1B = unname(params["m1B"]-params["m10"]),
m2A = unname(params["m2A"]-params["m20"]),
m2B = unname(params["m2B"]-params["m20"]), v=1)
param_plot <-
make_params_plot(params_to_plot, scale = 6) +
annotate("text", x = 1.375, y = 0.5,
label = paste0("v = ", params["v"]),
angle = -90, vjust = 1.5, size = 3)
panel_abund <-
out_pA_05 |>
as_tibble() |>
select(time, N1, N2) |>
pivot_longer(N1:N2) |>
ggplot(aes(x = time, y = value, color = name)) +
geom_line(linewidth = 0.9) +
scale_color_manual(values = c("#9970ab", "#5aae61"),
name = "Plant species", label = c("Plant 1", "Plant 2"), guide = "none") +
scale_y_continuous(breaks = c(0,8e17,1.6e18), labels = c("0", "7e17","1.4e18")) +
ylab("Abundance")
# Panel D: plot for frequencies of both plants when growing
# in dynamic soils
panel_freq <-
out_pA_05 |>
as_tibble() |>
mutate(p2 = 1-p1) |>
select(time, p1, p2) |>
pivot_longer(p1:p2) |>
ggplot(aes(x = time, y = value, color = name)) +
geom_line(linewidth = 1.2) +
scale_color_manual(values = c("#9970ab", "#5aae61"),
name = "Plant species", label = c("Plant 1", "Plant 2")) +
scale_y_continuous(limits = c(0,1), breaks = c(0, 0.5, 1)) +
ylab("Frequency")
param_plot + {panel_abund + panel_freq &
theme(axis.text = element_text(size = 8), legend.position = 'none')} +
plot_layout(widths = c(1/3, 2/3))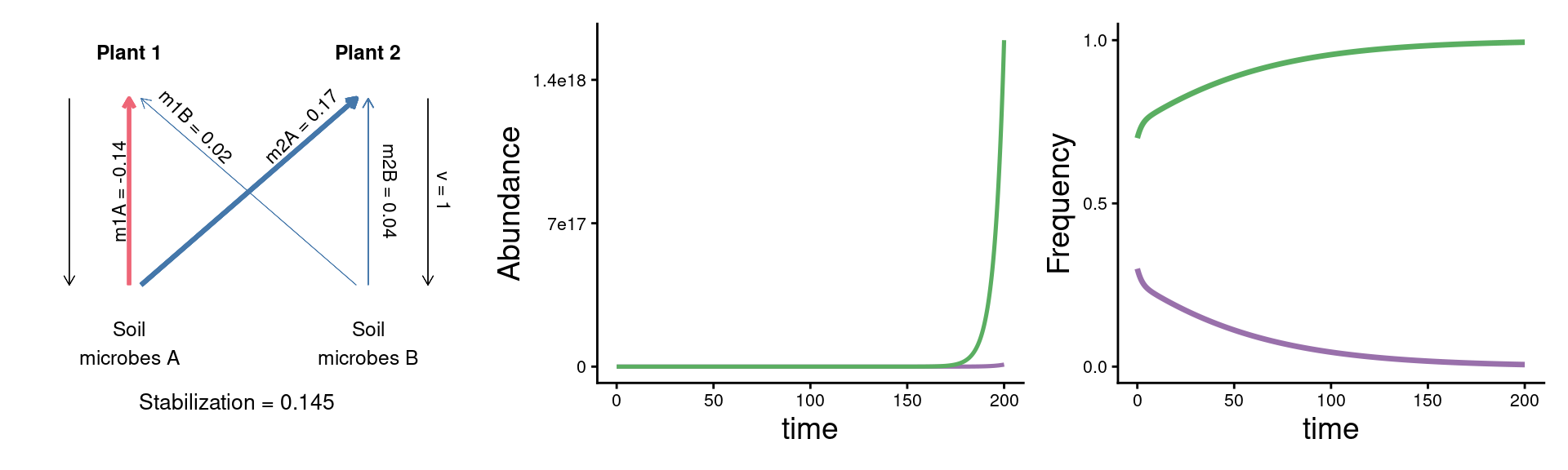
How can we more accurately infer predicted plant coexistence under plant–soil feedback?
Build on insights from coexistence theory (Chesson 2018):
Stabilization is necessary for coexistence but doesn’t guarantee coexistence
Stable coexistence is possible only when stabilization overcomes competitive imbalances (fitness differences)

How can we more accurately infer predicted plant dynamics under plant–soil feedback?
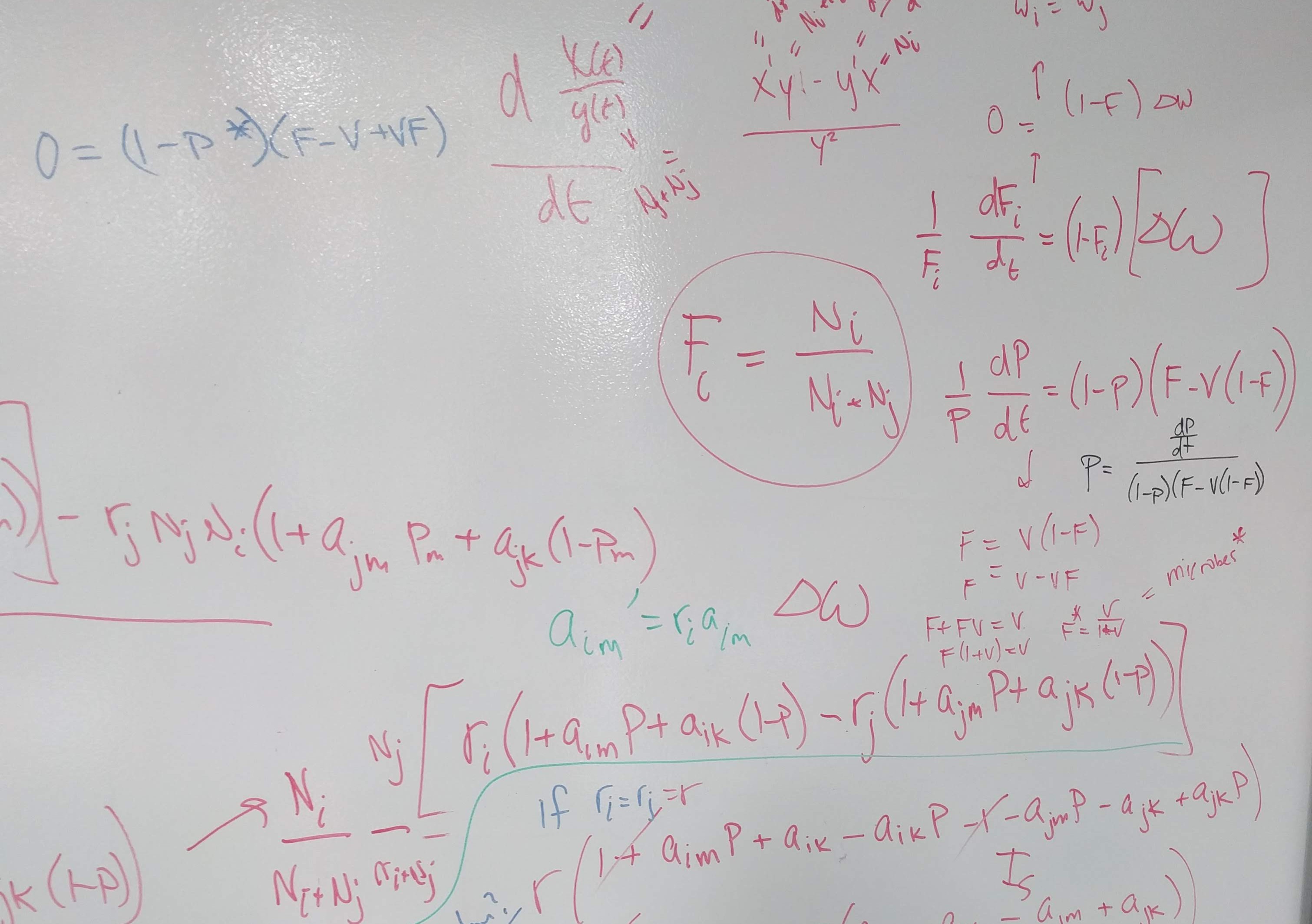
How can we more accurately infer predicted plant dynamics under plant–soil feedback?

How can we more accurately infer predicted plant dynamics under plant–soil feedback?

How can we more accurately infer predicted plant dynamics under plant–soil feedback?

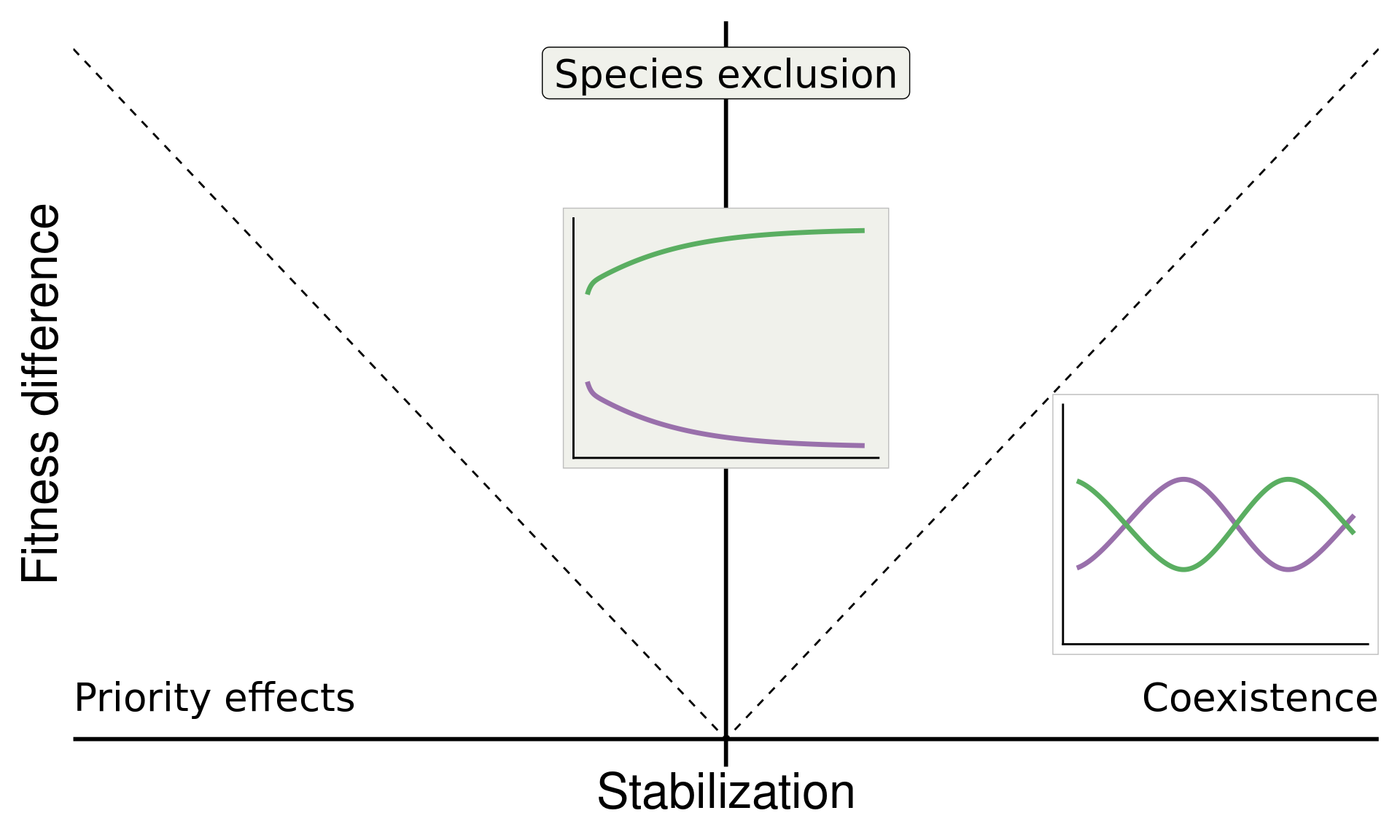
\[\text{Fitness difference}_{1,2} = \overbrace{\frac{1}{2}(m_{1A}+m_{1B})}^{\substack{\text{Sensitivity of}\\ \text{Sp 1 to microbes}}} - \overbrace{\frac{1}{2}(m_{2A}+m_{2B})}^{\substack{\text{Sensitivity of}\\ \text{Sp 2 to microbes}}}\]
Code
base <-
tibble(sd = seq(-1,1,0.1),
fd = c(seq(1,0,-0.1), seq(0.1,1,0.1))) |>
ggplot(aes(x = sd, y = fd)) +
geom_line(linetype = "dashed") +
geom_hline(yintercept = 0, linewidth = 1) +
geom_vline(xintercept = 0, linewidth = 1) +
xlab("Stabilization") +
ylab("Fitness difference") +
theme(axis.text = element_blank(),
axis.ticks = element_blank(),
axis.line = element_blank(),
axis.title = element_text(size = 25)) +
scale_x_continuous(expand = c(0,0)) +
scale_y_continuous(expand = c(0.04,0))
base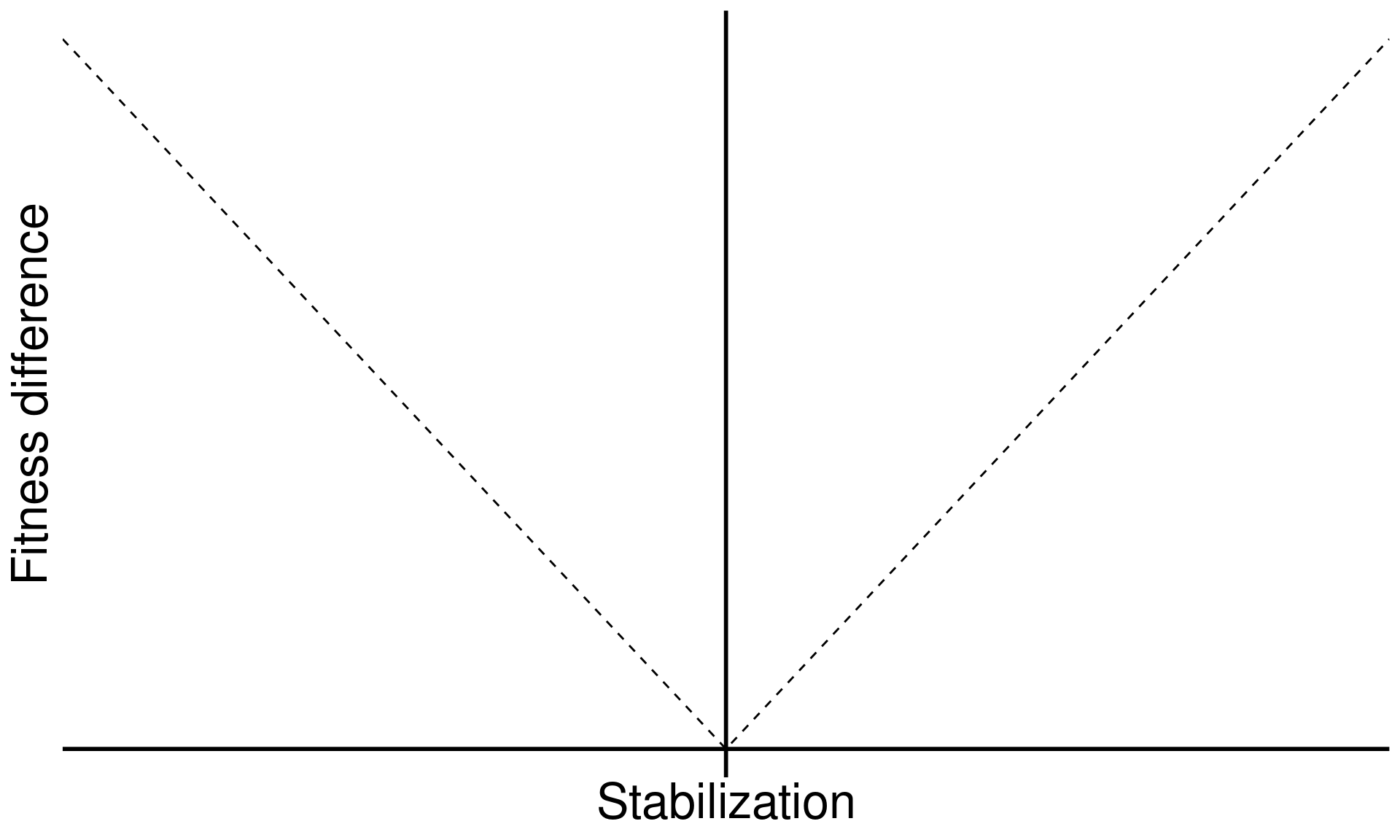
Code
base <-
base +
annotate("text", x = Inf, y = 0, hjust = 1, vjust = -1, label = "Coexistence", size = 7, family = "italic") +
annotate("text", x = -Inf, y = 0, hjust = 0, vjust = -1, label = "Priority effects", size = 7, family = "italic") +
annotate("label", x = 0, y = Inf, vjust = 1.5, label = "Species exclusion", size = 7, fill = "#F0F1EB",family = "italic")
base
Code
metrics <- function(params) {
IS <- with(as.list(params), {m1A - m2A - m1B + m2B})
FD <- with(as.list(params), {(1/2)*(m1A+m2A) - (1/2)*(m1B+m2B)})
SD <- (-1/2)*IS
return(c(IS = IS, SD = SD, FD = FD))
}
coex_metrics <- metrics(params_coex)
base_coex <-
base +
inset_element({panel_freq_coex + theme(legend.position = 'none',
axis.text = element_blank(),
axis.ticks = element_blank(),
axis.title = element_blank(),
plot.background = element_rect(color = "grey"))},
0.75,0.15,1,0.5)
base_coex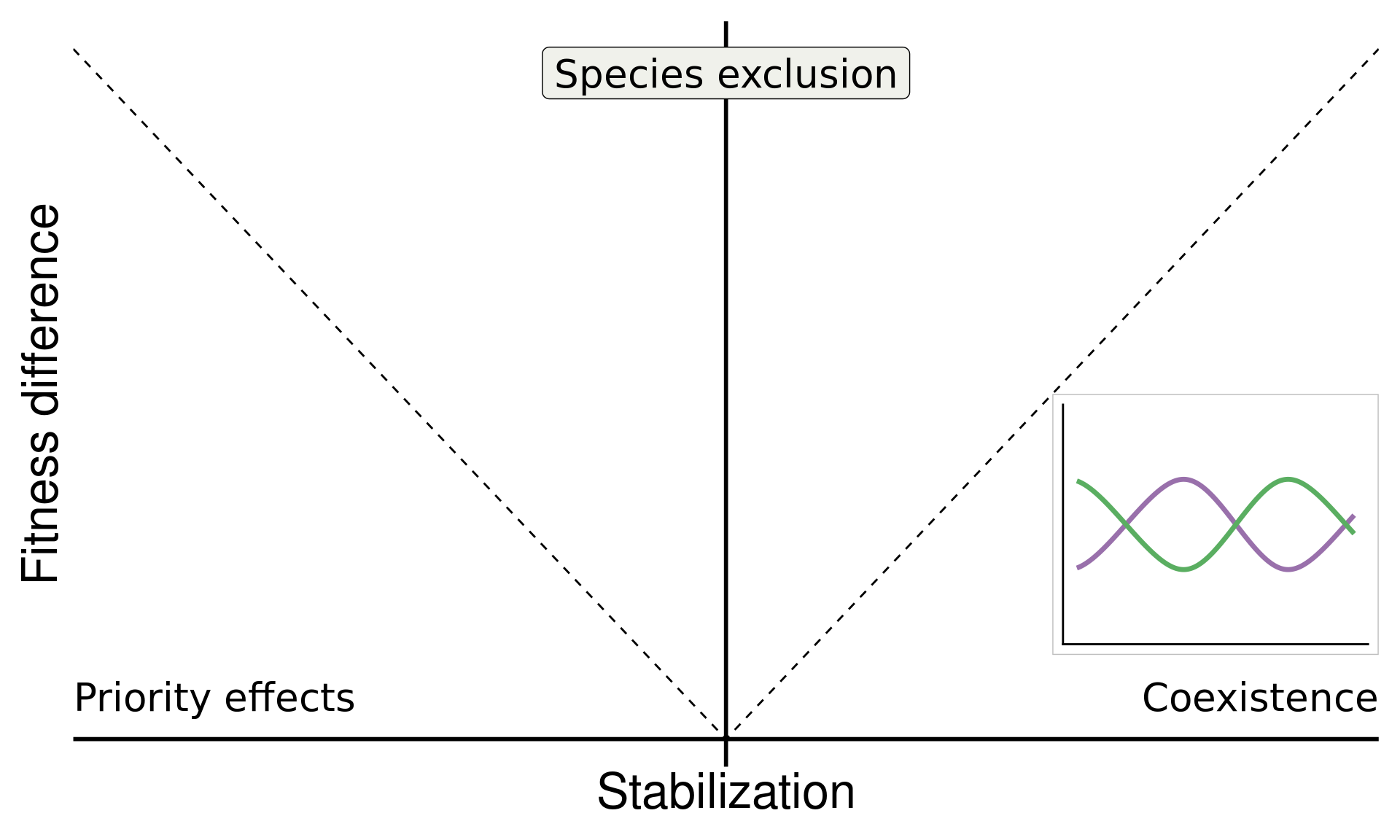
Do microbes generate fitness differences in nature?
Sedgwick Reserve (Santa Barbara County, California)
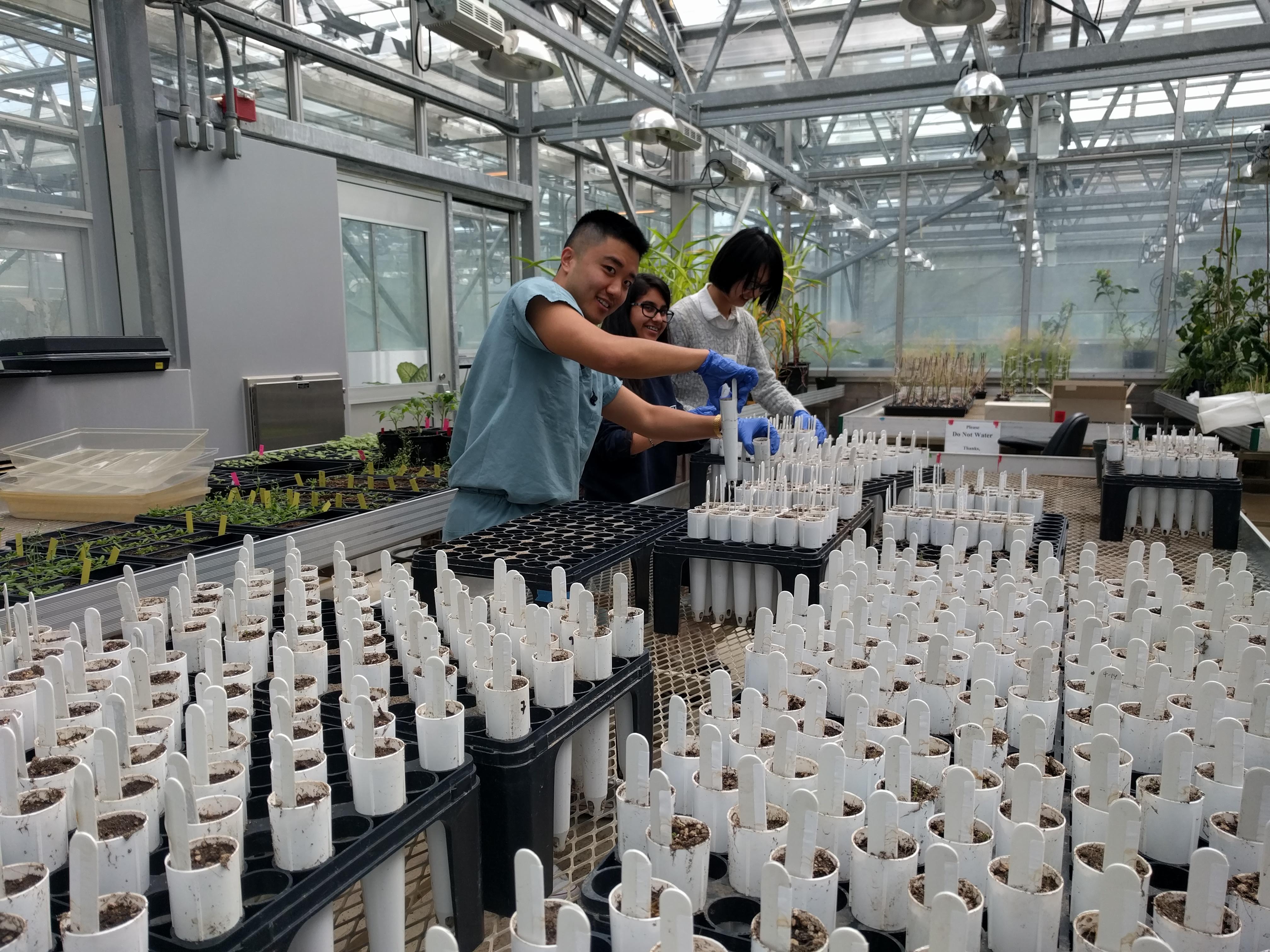

Code
# Dataset downloaded from Figshare:
# curl::curl_download("https://datadryad.org/downloads/file_stream/367188", destfile = "../data/kandlikar2021-supplement.csv")
biomass_wide <-
read_csv("../data/kandlikar2021-supplement.csv") |>
mutate(log_agb = log(abg_dry_g)) |>
mutate(source_soil = ifelse(source_soil == "aastr", "str", source_soil),
source_soil = ifelse(source_soil == "abfld", "fld", source_soil),
pair = paste0(source_soil, "_", focal_species)) |>
select(replicate, pair, log_agb) |>
pivot_wider(names_from = pair, values_from = log_agb) |>
filter(!is.na(replicate))
# Calculate the Stabilization between each species pair ----
# Recall that following the definition of the m terms in Bever 1997,
# and the analysis of this model in Kandlikar 2019,
# stablization = -0.5*(log(m1A) + log(m2B) - log(m1B) - log(m2A))
# Here is a function that does this calculation
# (recall that after the data reshaping above,
# each column represents a given value of log(m1A))
calculate_stabilization <- function(df) {
df |>
# In df, each row is one repilcate/rack, and each column
# represents the growth of one species in one soil type.
# e.g. "FE_ACWR" is the growth of FE in ACWR-cultivated soil.
mutate(AC_FE = -0.5*(AC_ACWR - AC_FEMI - FE_ACWR + FE_FEMI),
AC_HO = -0.5*(AC_ACWR - AC_HOMU - HO_ACWR + HO_HOMU),
AC_SA = -0.5*(AC_ACWR - AC_SACO - SA_ACWR + SA_SACO),
AC_PL = -0.5*(AC_ACWR - AC_PLER - PL_ACWR + PL_PLER),
AC_UR = -0.5*(AC_ACWR - AC_URLI - UR_ACWR + UR_URLI),
FE_HO = -0.5*(FE_FEMI - FE_HOMU - HO_FEMI + HO_HOMU),
FE_SA = -0.5*(FE_FEMI - FE_SACO - SA_FEMI + SA_SACO),
FE_PL = -0.5*(FE_FEMI - FE_PLER - PL_FEMI + PL_PLER),
FE_UR = -0.5*(FE_FEMI - FE_URLI - UR_FEMI + UR_URLI),
HO_PL = -0.5*(HO_HOMU - HO_PLER - PL_HOMU + PL_PLER),
HO_SA = -0.5*(HO_HOMU - HO_SACO - SA_HOMU + SA_SACO),
HO_UR = -0.5*(HO_HOMU - HO_URLI - UR_HOMU + UR_URLI),
SA_PL = -0.5*(SA_SACO - SA_PLER - PL_SACO + PL_PLER),
SA_UR = -0.5*(SA_SACO - SA_URLI - UR_SACO + UR_URLI),
PL_UR = -0.5*(PL_PLER - PL_URLI - UR_PLER + UR_URLI)) |>
select(replicate, AC_FE:PL_UR) |>
gather(pair, stabilization, AC_FE:PL_UR)
}
stabilization_values <- calculate_stabilization(biomass_wide)
# Seven values are NA; we can omit these
stabilization_values <- stabilization_values |>
filter(!(is.na(stabilization)))
# Calculating the Fitness difference between each species pair ----
# Similarly, we can now calculate the fitness difference between
# each pair. Recall that FD = 0.5*(log(m1A)+log(m1B)-log(m2A)-log(m2B))
# But recall that here, IT IS IMPORTANT THAT
# m1A = (m1_soilA - m1_fieldSoil)!
# The following function does this calculation:
calculate_fitdiffs <- function(df) {
df |>
mutate(AC_FE = 0.5*((AC_ACWR-fld_ACWR) + (FE_ACWR-fld_ACWR) - (AC_FEMI-fld_FEMI) - (FE_FEMI-fld_FEMI)),
AC_HO = 0.5*((AC_ACWR-fld_ACWR) + (HO_ACWR-fld_ACWR) - (AC_HOMU-fld_HOMU) - (HO_HOMU-fld_HOMU)),
AC_SA = 0.5*((AC_ACWR-fld_ACWR) + (SA_ACWR-fld_ACWR) - (AC_SACO-fld_SACO) - (SA_SACO-fld_SACO)),
AC_PL = 0.5*((AC_ACWR-fld_ACWR) + (PL_ACWR-fld_ACWR) - (AC_PLER-fld_PLER) - (PL_PLER-fld_PLER)),
AC_UR = 0.5*((AC_ACWR-fld_ACWR) + (UR_ACWR-fld_ACWR) - (AC_URLI-fld_URLI) - (UR_URLI-fld_URLI)),
FE_HO = 0.5*((FE_FEMI-fld_FEMI) + (HO_FEMI-fld_FEMI) - (FE_HOMU-fld_HOMU) - (HO_HOMU-fld_HOMU)),
FE_SA = 0.5*((FE_FEMI-fld_FEMI) + (SA_FEMI-fld_FEMI) - (FE_SACO-fld_SACO) - (SA_SACO-fld_SACO)),
FE_PL = 0.5*((FE_FEMI-fld_FEMI) + (PL_FEMI-fld_FEMI) - (FE_PLER-fld_PLER) - (PL_PLER-fld_PLER)),
FE_UR = 0.5*((FE_FEMI-fld_FEMI) + (UR_FEMI-fld_FEMI) - (FE_URLI-fld_URLI) - (UR_URLI-fld_URLI)),
HO_PL = 0.5*((HO_HOMU-fld_HOMU) + (PL_HOMU-fld_HOMU) - (HO_PLER-fld_PLER) - (PL_PLER-fld_PLER)),
HO_SA = 0.5*((HO_HOMU-fld_HOMU) + (SA_HOMU-fld_HOMU) - (HO_SACO-fld_SACO) - (SA_SACO-fld_SACO)),
HO_UR = 0.5*((HO_HOMU-fld_HOMU) + (UR_HOMU-fld_HOMU) - (HO_URLI-fld_URLI) - (UR_URLI-fld_URLI)),
SA_PL = 0.5*((SA_SACO-fld_SACO) + (PL_SACO-fld_SACO) - (SA_PLER-fld_PLER) - (PL_PLER-fld_PLER)),
SA_UR = 0.5*((SA_SACO-fld_SACO) + (UR_SACO-fld_SACO) - (SA_URLI-fld_URLI) - (UR_URLI-fld_URLI)),
PL_UR = 0.5*((PL_PLER-fld_PLER) + (UR_PLER-fld_PLER) - (PL_URLI-fld_URLI) - (UR_URLI-fld_URLI))) |>
select(replicate, AC_FE:PL_UR) |>
gather(pair, fitdiff_fld, AC_FE:PL_UR)
}
fd_values <- calculate_fitdiffs(biomass_wide)
# Twenty-two values are NA; let's omit these.
fd_values <- fd_values |> filter(!(is.na(fitdiff_fld)))
# Now, generate statistical summaries of SD and FD
stabiliation_summary <- stabilization_values |> group_by(pair) |>
summarize(mean_sd = mean(stabilization),
sem_sd = sd(stabilization)/sqrt(n()),
n_sd = n())
fitdiff_summary <- fd_values |> group_by(pair) |>
summarize(mean_fd = mean(fitdiff_fld),
sem_fd = sd(fitdiff_fld)/sqrt(n()),
n_fd = n())
# Combine the two separate data frames.
sd_fd_summary <- left_join(stabiliation_summary, fitdiff_summary)
# Some of the FDs are negative, let's flip these to be positive
# and also flip the label so that the first species in the name
# is always the fitness superior.
sd_fd_summary <- sd_fd_summary |>
# if mean_fd is < 0, the following command gets the absolute
# value and also flips around the species code so that
# the fitness superior is always the first species in the code
mutate(pair = ifelse(mean_fd < 0,
paste0(str_extract(pair, "..$"),
"_",
str_extract(pair, "^..")),
pair),
mean_fd = abs(mean_fd))
sd_fd_summary <-
sd_fd_summary |>
mutate(
# outcome = ifelse(mean_fd - 2*sem_fd >
# mean_sd + 2*sem_sd, "exclusion", "neutral"),
outcome2 = ifelse(mean_fd > mean_sd, "exclusion", "coexistence"),
outcome2 = ifelse(mean_sd < 0, "exclusion or priority effect", outcome2)
)
base_w_rug <-
base +
geom_point(data = sd_fd_summary,
aes(x = mean_sd, y = 0), shape = "|", size = 5,
inherit.aes = F) +
theme(axis.title = element_blank())
base_w_rug
Code
# Example point, using UR_FE values
base_w_rug +
annotate("pointrange", x = 0.275, y = 0.0591, ymin = 0.0591-0.307,ymax = 0.0591+0.307, shape = 21, stroke = 1.5, size = 1, linewidth = 0.1, fill = "#4477aa") +
annotate("pointrange", x = 0.275, y = 0.0591, xmin = 0.275-0.152,xmax = 0.275+0.152, shape = 21, fill = "#4477aa", linewidth = 0.1, size = 1, stroke = 1.5) +
annotate("text", x = 0.275 + 0.3, y = 0.0591 + 0.15, color = "#4477aa", label = "Festuca - Plantago", size = 8, fontface = "italic") 
Code
base +
geom_pointrange(data = sd_fd_summary,
aes(x = mean_sd, y = mean_fd,
ymin = mean_fd-sem_fd,
ymax = mean_fd+sem_fd, fill = outcome2), size = 1, linewidth = 0.1,
shape = 21, stroke = 1.5) +
geom_pointrange(data = sd_fd_summary,
aes(x = mean_sd, y = mean_fd,
xmin = mean_sd-sem_sd,
xmax = mean_sd+sem_sd, fill = outcome2), size = 1, linewidth = 0.1, shape = 21, stroke = 1.5) +
scale_fill_manual(values = c("#4477aa", "#ee6677", "#ccbb44")) +
theme(legend.position = "none")+
theme(axis.title = element_blank())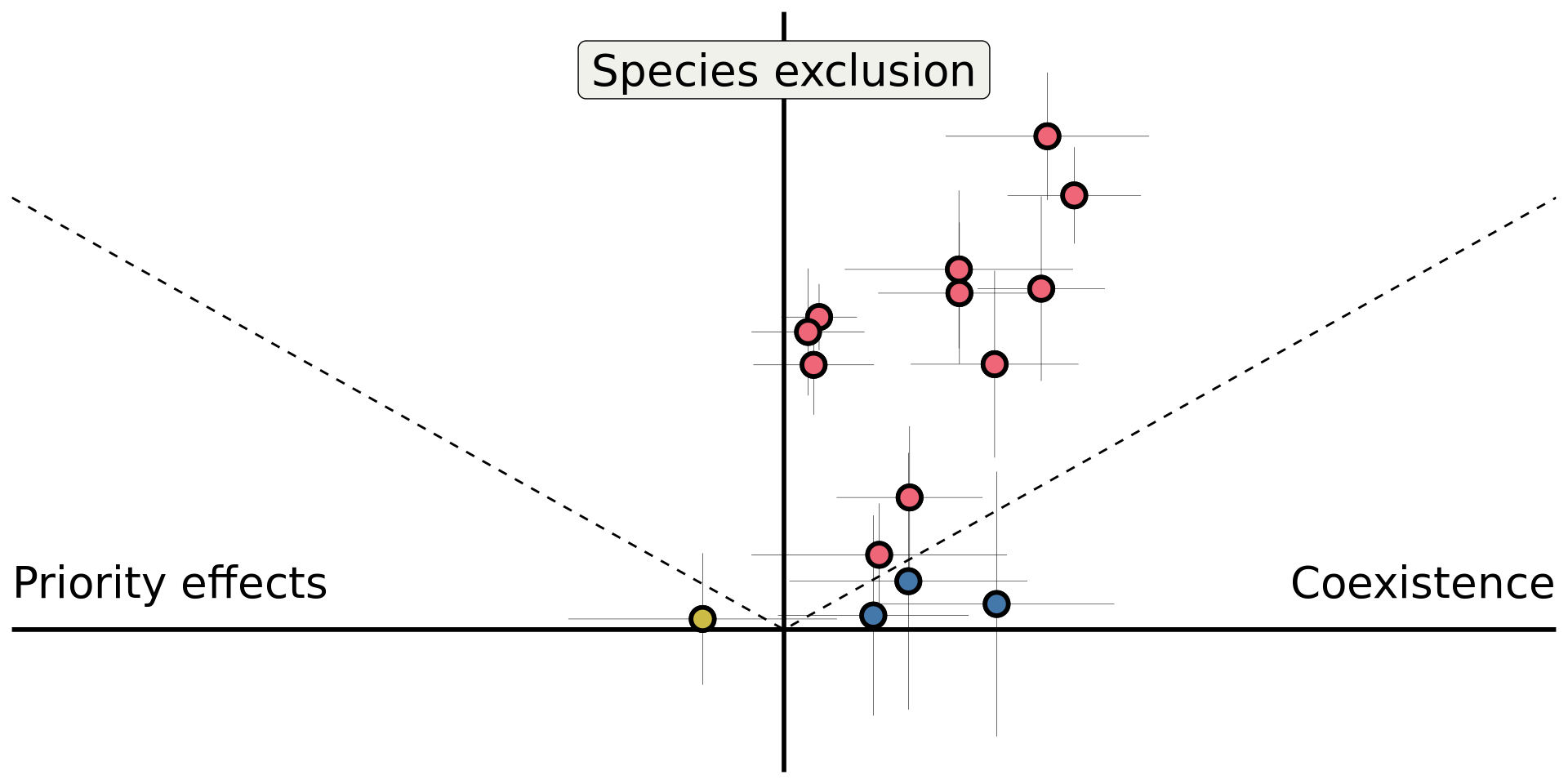
Detailed in Kandlikar et al. (2021)
Do microbes generate fitness differences in nature?
In California grasslands, microbes tend to drive stronger fitness differences than stabilization.
Can we answer this question more generally?

Meta-analysis overview
Data for 518 pairwise comparisons of stabilization and fitness differences (results shown for 72 pairs here)
Results in Yan, Levine, and Kandlikar (2022); Analyses available on Zenodo
Do microbes generate fitness differences in nature?
- Fitness differences represent a dominant avenue through which microbes control plant coexistence
Microbially-mediated coexistence in pine-oak forests in Spain, Pajares-Murgó et al. (2024)
Unceded Chumash territory
Today’s talk
- Ecological theory for microbial effects on plant coexistence
2. Microbial effects on plant communities in variable environments
General themes:
Advancing ecological insights in light of conservation and management challenges
Situating our understanding in historical contexts
Plant–soil–fire impacts on woody plant encroachment
Historical range of the longleaf pine ecosystem
Code
library(sf)
library(tidyverse)
little <- sf::st_read("../img/pinupalu.shp")
## Reading layer `pinupalu' from data source
## `/home/gkandlikar@lsu.edu/gklab/talks/img/pinupalu.shp' using driver `ESRI Shapefile'
## Simple feature collection with 38 features and 5 fields
## Geometry type: POLYGON
## Dimension: XY
## Bounding box: xmin: -95.21806 ymin: 26.61921 xmax: -75.79999 ymax: 36.84856
## CRS: NA
# States we want to show
llp_states <- c("texas", "louisiana","mississippi", "alabama", "florida", "georgia", "south carolina", "north carolina", "virginia", "tennessee")
states <- map_data("state") |>
filter(region %in% llp_states)
ggplot() +
geom_polygon(data = states,
aes(x = long, y = lat, group = group), fill = "transparent", color = 'darkgrey') +
geom_sf(data = little, fill = alpha('#228833', 0.5)) +
theme_void()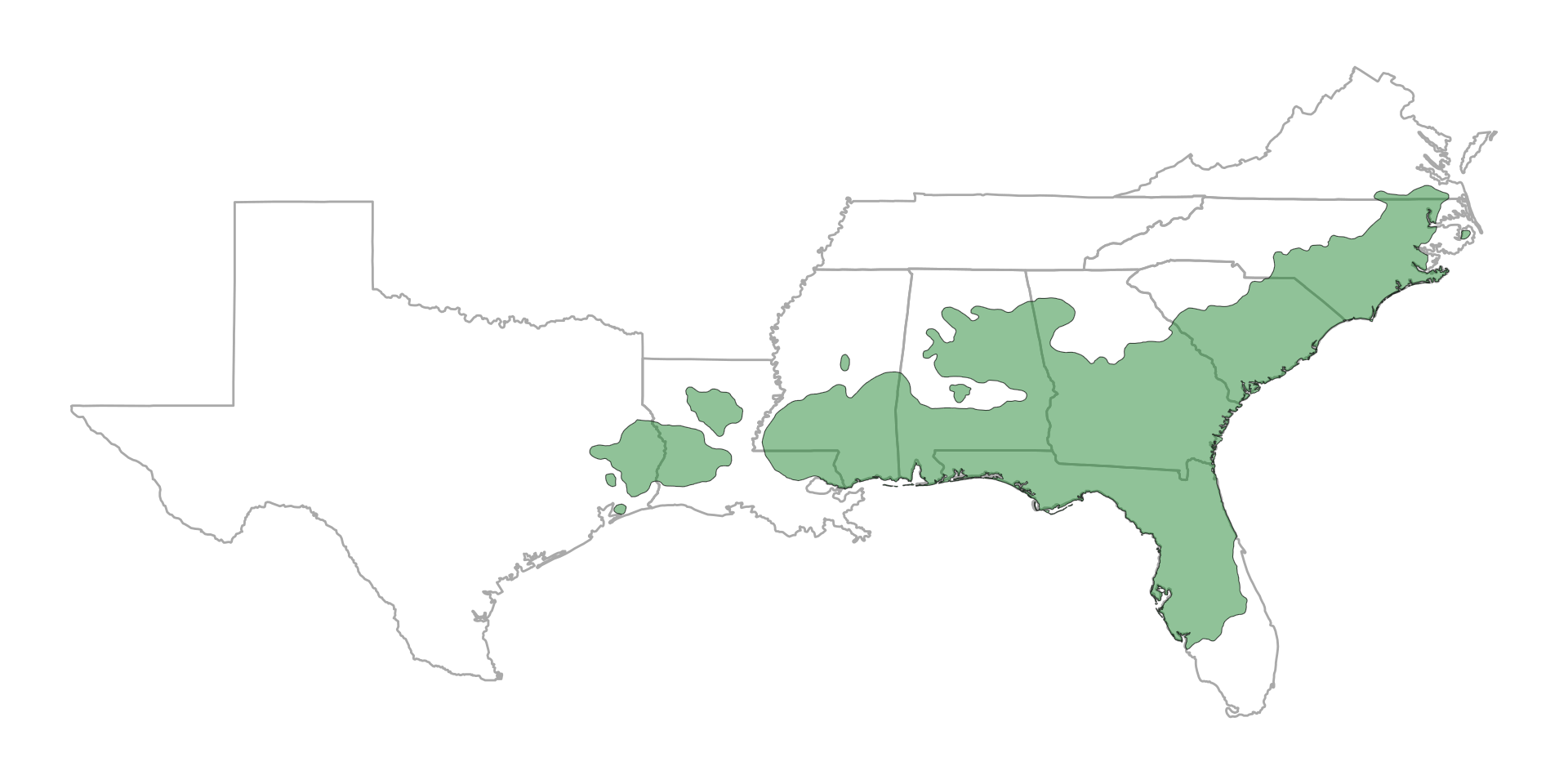
Photo: Kandlikar lab members at Lee Memorial Forest (LSU Ag Center), Franklinton LA
Prescribed burn at LSU Forestry Camp, 13 April 2026
Turpentine Industry from Frank Leslie’s Illustrated Newspaper. Link.
Photo from Longleaf Alliance. Link.
Still image from 1920s US Forest Service film “When Trees Talk”. Video available here.
Fire helps maintain the open understory


Remarkable diversity at small spatial scales (40-50 plant spp/m2)
Woody encroachment pervasive under fire suppression
Woody encroachment is common worldwide
Over 500 million hectares of grasslands/savannas affected
Many drivers…
Woody encroachment is common worldwide
Over 500 million hectares of grasslands/savannas affected
Many drivers… but we wonder if there is a role for plant–soil microbial feedbacks.

Might microbes play a role?
Two lines of evidence.
1. Across ecosystems, grasses tend to grow worse with conspecific-conditioned soil microbes
Code
# curl::curl_download("https://datadryad.org/downloads/file_stream/2789266", destfile = "../data/jiang2024-dataset.csv")
jiang_dat <- read_csv("../data/jiang2024-dataset.csv")
# jiang_metamod <- metafor::rma.mv(rr, var, data = jiang_dat, mods = ~ Life.form-1)
# saveRDS(jiang_metamod, "../data/jiang-metamod.rds")
metamod <- readRDS("../data/jiang-metamod.rds")
metamod_s <- broom::tidy(metamod)
jiang_plot <-
jiang_dat |>
ggplot(aes(x = rr, y = Life.form, color = Life.form)) +
ggbeeswarm::geom_quasirandom(shape = 21) +
scale_color_manual(values = c("#ccbb44","transparent")) +
annotate("point",
x = metamod_s$estimate[1], y = 1, size = 3, shape = 21, stroke = 1.5) +
# annotate("point",
# x = metamod_s$estimate[2], y = 2, size = 3, shape = 21, stroke = 1.5) +
ylab("") +
xlab("Growth in self vs. growth in non-self") +
xlim(-2.5,2.5)+
geom_vline(xintercept = 0, linetype = 'dashed') +
theme(legend.position = "none",
axis.text.y = element_text(size = 12, color = 'black'))
jiang_plot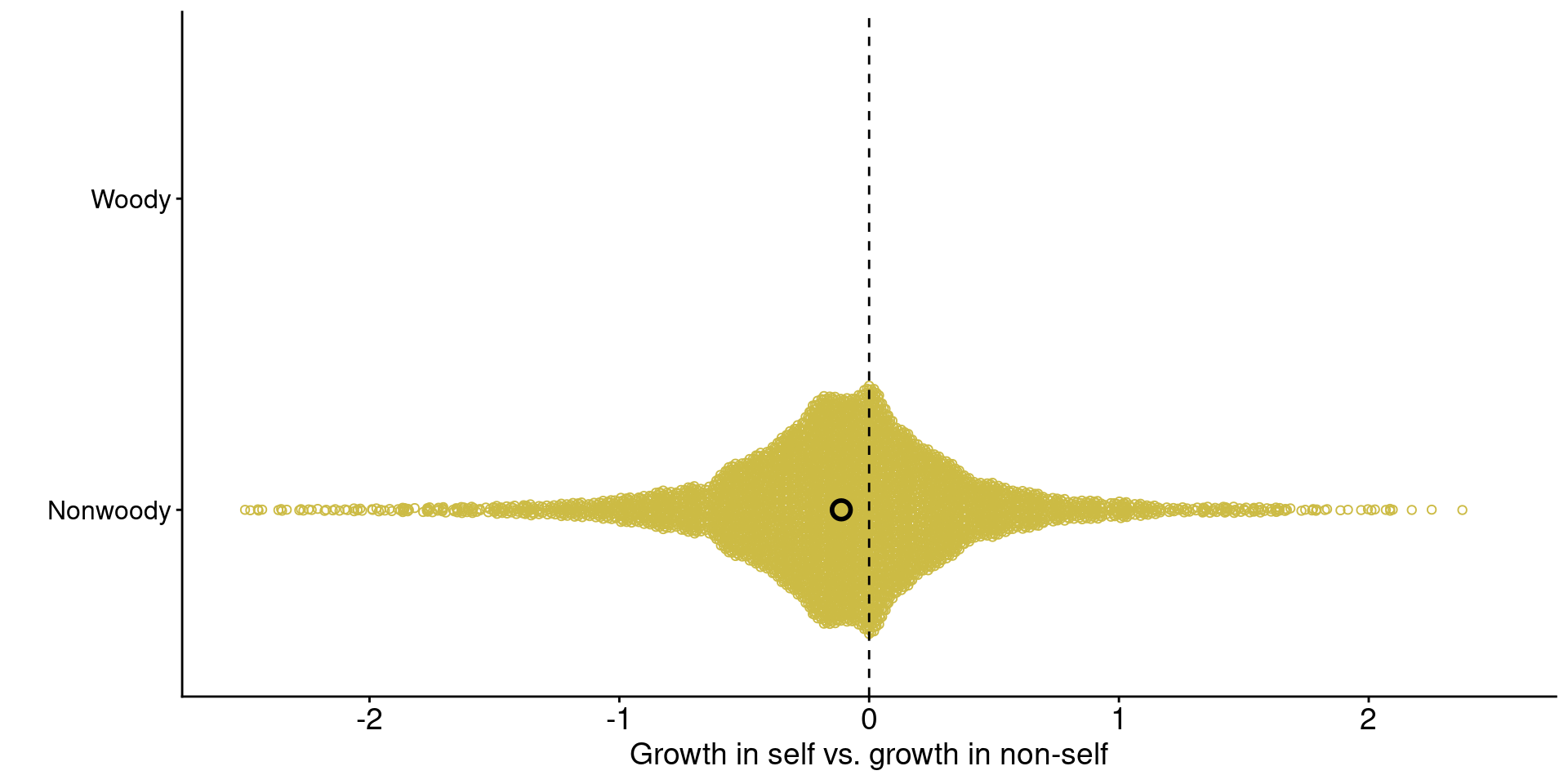
Data from Jiang et al. (2024)
1. Across ecosystems, grasses tend to grow worse with conspecific-conditioned soil microbes
Code
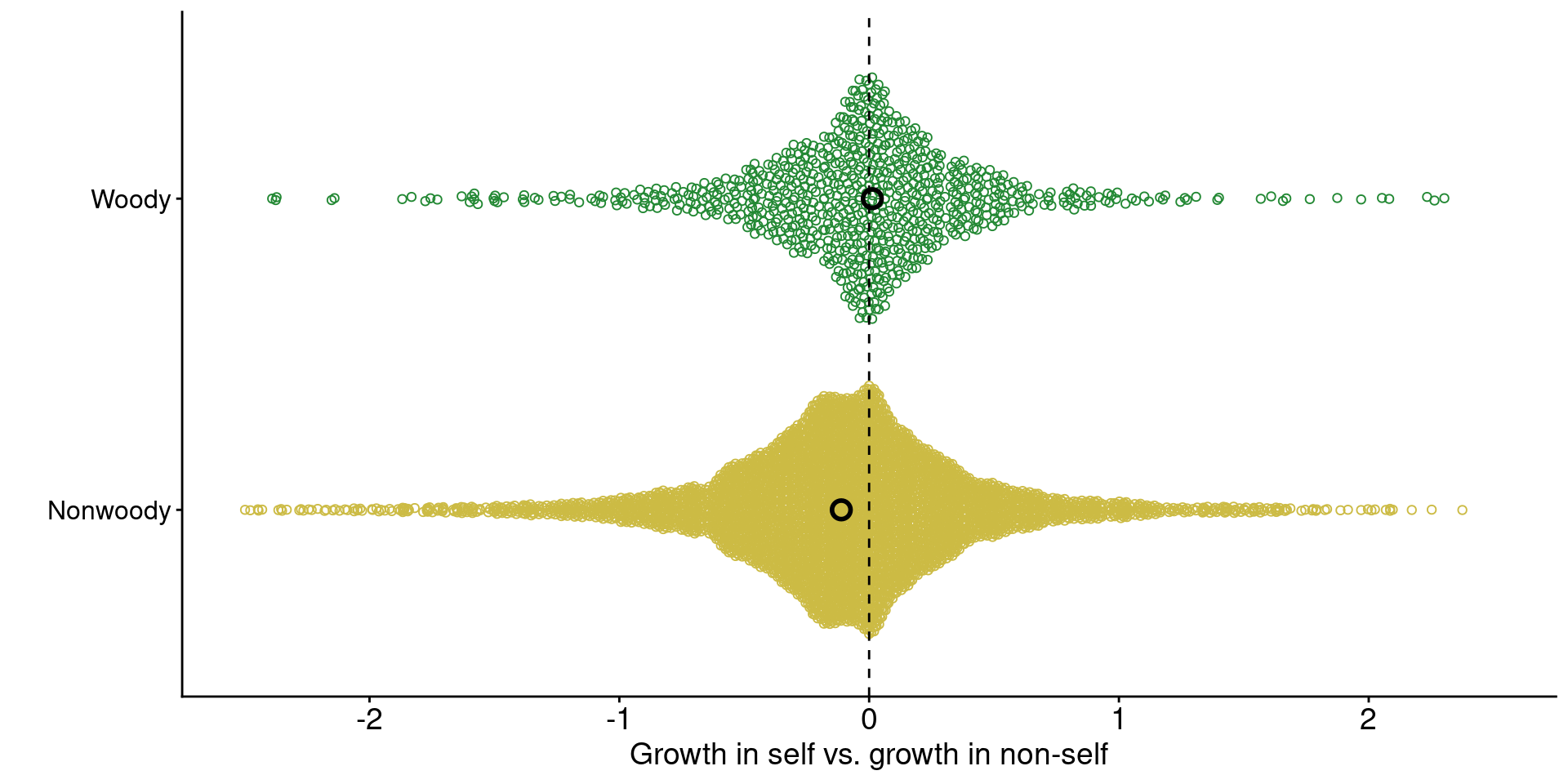
Data from Jiang et al. (2024)
2. Encroaching woody plants often have distinct microbial symbionts.
In African savannas, N-fixing legumes are the primary woody invader in >90% of studies sites (Stevens et al. 2017)
Ectomycorrhizal symbiosis is especially more common among encroaching woody plants (Simha and Kandlikar 2026)
Code
# download.file("https://www.science.org/doi/suppl/10.1126/science.aai8212/suppl_file/bennett_aai8212_database-s1.xlsx", destfile = "../data/bennett2017-dataset.xlsx")
readxl::read_xlsx("../data/bennett-2017.xlsx", sheet = 2) |>
mutate(`Type of Mycorrhiza` = factor(`Type of Mycorrhiza`, c("EM", "AM"))) |>
mutate(logratio = log(`Average of Biomass in Conspecific Soil`/`Average of Biomass in Heterospecific Soil`)) |>
arrange(logratio) |>
mutate(rown = row_number()) |>
ggplot(aes(x = logratio, color = `Type of Mycorrhiza`, y = rown, shape = `Type of Mycorrhiza`)) +
geom_point(size = 2) +
geom_segment(aes(x = 0, xend = logratio), linewidth = 0.125) +
xlab("Growth in self-conditioned microbes vs.\n nonself-conditioned microbes") +
geom_vline(xintercept = 0) +
scale_color_manual(values = c("#0077bb", "#cc3311")) +
theme(axis.line.y = element_blank(),
axis.text.y = element_blank(),
axis.title.y = element_blank(),
axis.ticks.y = element_blank(),
legend.position = "inside",
legend.position.inside = c(0.9, 0.1),
legend.text = element_text(size = 12),
legend.title = element_blank())
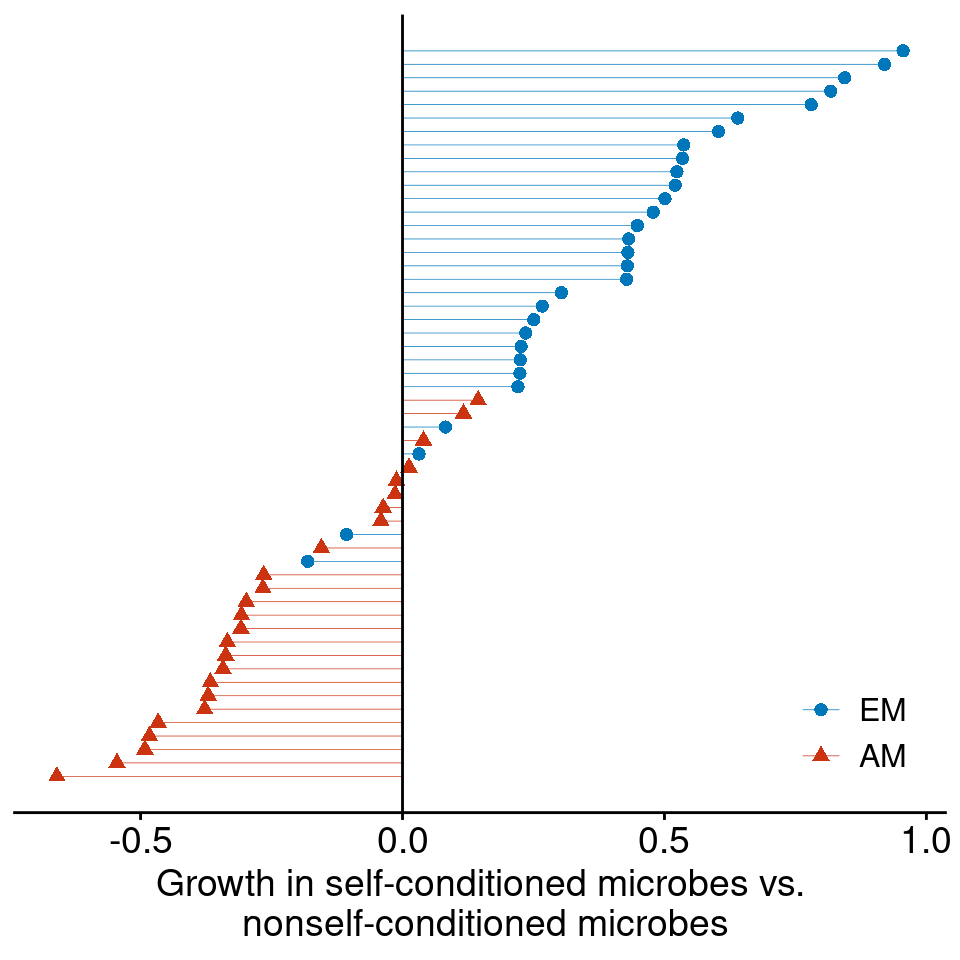
Data from Bennett et al. (2017)
Do soil microbes play a role in driving woody encroachment under fire suppression?
Approach:
Mathematical modeling: experimentally-informed dynamics of plant community dynamics
Field observations and experiment: evaluate fire effects on soil communities
Plant growth experiment: evaluate shrub and grass growth in pre- and post-fire shrub- and grass-conditioned soils
Mathematical modeling of plant dynamics
Transitions between patches
Patches without plants or soil conditioning
\[ \frac{dP_{00}}{dt} = - \overbrace{r_s P_{00}(P_{ss})}^{\substack{\text{woody} \\ \text{establishment}}}- \overbrace{r_g P_{00}(P_{{g0}+}P_{gg}+P_{gs})}^{\text{grass establishment}} + \overbrace{\mu_g P_{g0} +\mu_\sigma P_{\sigma 0}}^{\substack{\text{mortality in} \\ \text{unconditioned patch}}} + \overbrace{d_g P_{0g} + d_s P_{0s}}^{\substack{\text{microbial decay in} \\ \text{unoccupied patch}}} \qquad(1)\]
Patches with plant present but soil not yet conditioned
\[ \frac{dP_{g0}}{dt} = \overbrace{r_g P_{00}(P_{g0} + P_{gg} + P_{gs})}^{\substack{\text{grass establishment in} \\ \text{unconditioned patch}}} - \overbrace{\mu_g P_{g0}}^{\substack{\text{grass} \\ \text{mortality}}} - \overbrace{c_g P_{g0}} ^{\substack{\text{grass} \\ \text{conditioning}}} \qquad(2)\]
\[ \frac{dP_{\sigma 0}}{dt} = \overbrace{r_s P_{00}(P_{ss})}^{\substack{\text{woody establishment in} \\ \text{unconditioned patch}}} - \overbrace{\mu_\sigma P_{\sigma 0}}^{\substack{\text{woody} \\ \text{mortality}}} - \overbrace{c_s P_{\sigma 0}}^{\substack{\text{woody} \\ \text{conditioning}}} \qquad(3)\]Patches with plants and corresponding soil conditioning effects
\[ \frac{dP_{gg}}{dt} = \overbrace{r_g m_{gg} P_{0g}(P_{g0} + P_{gg} + P_{gs})}^{\substack{\text{grass establishment in} \\ \text{grass-conditioned patch}}} + \overbrace{c_g (P_{g0} + P_{gs})}^{\text{conditioning}} - \overbrace{\mu_g P_{gg}}^{\text{mortality}} \qquad(4)\]
\[ \frac{dP_{\sigma s}}{dt} = \overbrace{r_s m_{ss} P_{0s}P_{ss}}^{\substack{\text{woody establishment in} \\ \text{woody-conditioned patch}}} - \overbrace{c_s P_{\sigma s}}^{\substack{\text{growth into} \\ \text{adult stage}}} - \overbrace{\mu_\sigma P_{\sigma s}}^{\text{mortality}} \qquad(5)\]
\[ \frac{dP_{ss}}{dt} = \overbrace{c_s (P_{\sigma 0} + P_{\sigma g})}^{\substack{\text{conditioning by and growth of}\\ \text{woody seedlings} }} + \overbrace{c_s (P_{\sigma s})}^{\substack{\text{ growth of}\\ \text{woody seedlings} }} - \overbrace{\mu_s P_{ss}}^{\text{mortality}} \qquad(6)\]
Patches with plants growing in the other plant’s conditioned soil
\[ \frac{dP_{gs}}{dt} = \overbrace{r_g m_{gs} P_{0s} (P_{g0} + P_{gg} + P_{gs})}^{\substack {\text{grass establishment in} \\ \text{woody-conditioned patch}}} - \overbrace{\mu_g P_{gs}}^{\text{mortality}} - \overbrace{c_g P_{gs}}^{\text{conditioning}} \qquad(7)\]
\[ \frac{dP_{\sigma g}}{dt} = \overbrace{r_s m_{sg} P_{0g} (P_{ss})}^{\substack {\text{woody establishment in} \\ \text{grass-conditioned patch}}} - \overbrace{\mu_\sigma P_{\sigma g}}^{\text{mortality}} - \overbrace{c_s P_{\sigma g}}^{\text{conditioning}} \qquad(8)\]Patches with no plant present but conditioned soil persists
\[ \frac{dP_{0g}}{dt} = - \overbrace{r_g m_{gg} P_{0g}(P_{g0}+P_{gg}+P_{gs})}^{\text{grass establishment}} - \overbrace{r_s m_{sg} P_{0g}(P_{ss})}^{\substack{\text{woody} \\ \text{establishment}}} + \overbrace{\mu_g P_{gg} + \mu_\sigma P_{\sigma g}}^{\substack{\text{mortality in} \\ \text{grass-conditioned patch}}} - \overbrace{d_g P_{0g}}^{\substack{\text{microbial} \\ \text{decay}}} \qquad(9)\]
\[ \begin{multline} \frac{dP_{0s}}{dt} = - \overbrace{r_s m_{ss} P_{0s}(P_{ss})}^{\text{woody establishment}} - \overbrace{r_g m_{gs} P_{0s}(P_{g0}+P_{gg}+P_{gs})}^{\text{grass establishment}} + \\ \overbrace{\mu_s P_{ss} + \mu_\sigma P_{\sigma s} + \mu_g P_{gs}}^{\substack{\text{mortality in} \\ \text{woody-conditioned patch}}} - \overbrace{d_s P_{0s}}^{\substack{\text{microbial} \\ \text{decay}}} \end{multline} \qquad(10)\]Model dynamics in the absence of fire
Model dynamics in the absence of fire
Model dynamics in the absence of fire
Code
library(tidybayes)
meta <- readRDS("../img/swj-brms-meta.RDS")
plot_det <-
readRDS("../img/swj-plotdet.rds") +
labs(tag = "",
caption = "Note that priority effects can arise in the top-right quadrant.") +
theme(plot.caption = element_text(famile = "italic"))
b_draws <-
meta |>
spread_draws(b_Life.formNonwoody, b_Life.formWoody) |>
rename(mgg_draws = b_Life.formNonwoody,
mss_draws = b_Life.formWoody) |>
median_qi() |>
mutate(across(is.numeric, exp))
plot_det_pt <-
plot_det +
geom_pointrange(inherit.aes = F,
data = b_draws,
aes(x = mgg_draws, y = mss_draws,
xmin = mgg_draws.lower, xmax = mgg_draws.upper),
color = 'white', shape = 21, size = 1, stroke = 2) +
geom_pointrange(inherit.aes = F,
data = b_draws,
aes(x = mgg_draws, y = mss_draws,
ymin = mss_draws.lower, ymax = mss_draws.upper),
color = 'white', shape = 21, size = 1, stroke = 2) +
labs(caption = "White point & error bars show median & 95 CrI of Bayesian meta-analysis\nof grasses and woody plants from areas undergoing encroachment.")
plot_det_pt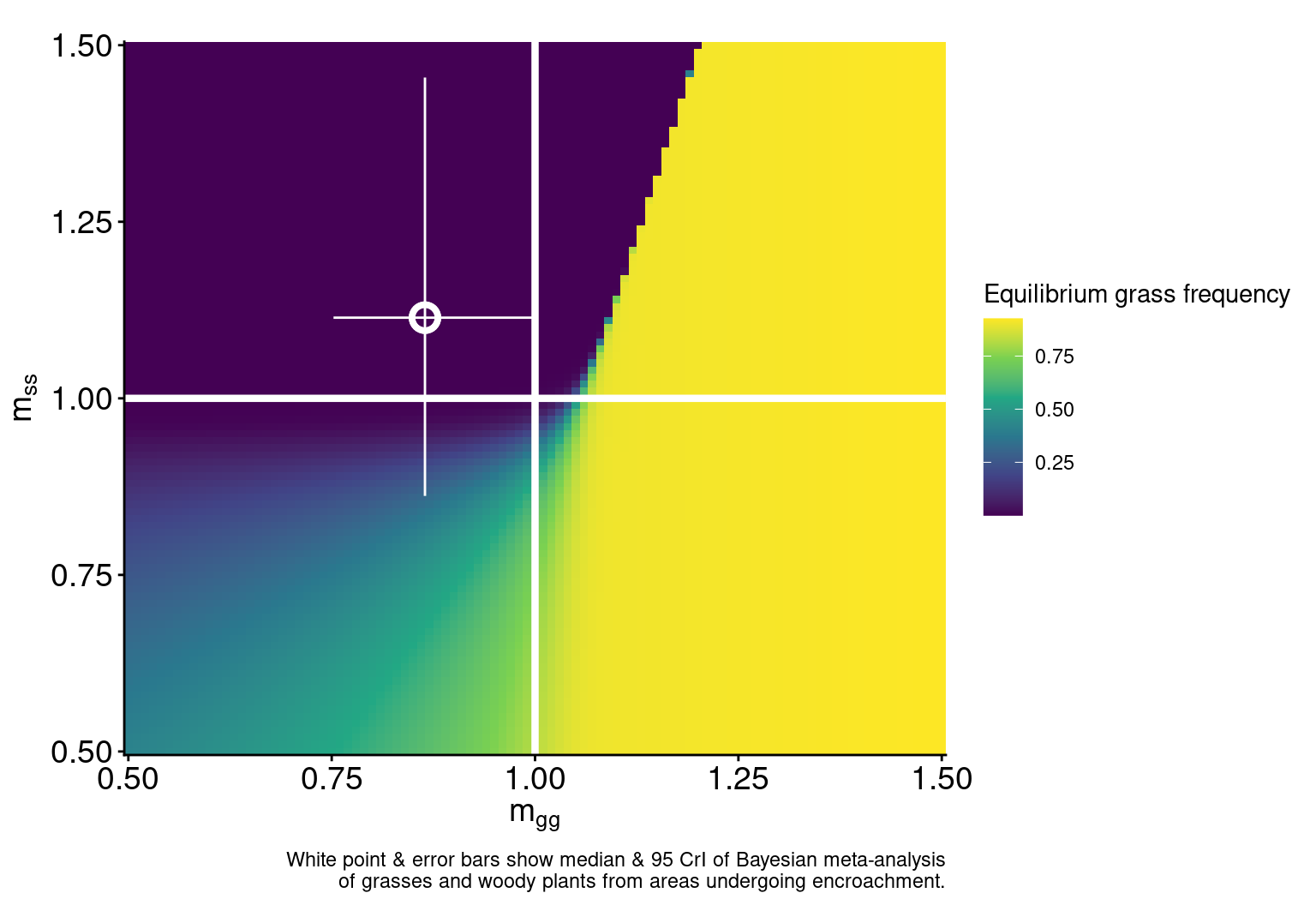
Fires alter plant and microbial states
Fires alter plant and microbial states
Fires of higher intensities cause more mortality of grasses, woody seedlings, and soil microbes.
Three scenarios of fire impacts on woody adults:
Code
int = seq(0,1, length.out = 20)
fire = data.frame(int = seq(0,1, length.out = 20),
grass = 1/(1+150*exp(-10.02*int)),
seedling = 1/(1+150*exp(-20.04*int)),
adult = 1/(1+150*exp(-10.02*int)),
sens = rep("Fire Sensitive", 20)) |>
pivot_longer(cols=c(grass,seedling,adult), names_to = "Plant", values_to = "mort")
fire2 = fire |> mutate(sens = "Less Sensitive",
mort = case_when(Plant == "adult" ~ mort/2, TRUE ~ mort))
fire3 = fire |> mutate(sens = "Not Sensitive",
mort = case_when(Plant == "adult" ~ 0, TRUE ~ mort))
fire_all = rbind(fire, fire2, fire3) |> mutate(sens = as.factor(sens), Plant = as.factor(Plant))
levels(fire_all$sens)
## [1] "Fire Sensitive" "Less Sensitive" "Not Sensitive"
levels(fire_all$Plant) = c("Woody Adult", "Grass", "Woody Seedling")
fire_all$Plant = factor(fire_all$Plant, levels = c("Grass", "Woody Seedling", "Woody Adult"))
scenarios = ggplot(data = fire_all, aes(x = int, y = mort, group = Plant)) +
geom_path(aes(color = Plant, linetype = Plant), linewidth = 3) +
scale_color_manual(values = c("#FDE725FF", "#2A788EFF", "#440154FF"))+
scale_linetype_manual(values = c("solid", "dashed", "longdash"))+
labs(x = "Fire Intensity", y="Percent Mortality")+
scale_x_continuous(expand = c(0,0), breaks = c(0.25, 0.5, 0.75, 1.0)) +
scale_y_continuous(expand = c(0,0))+
facet_wrap(~ sens)
scenarios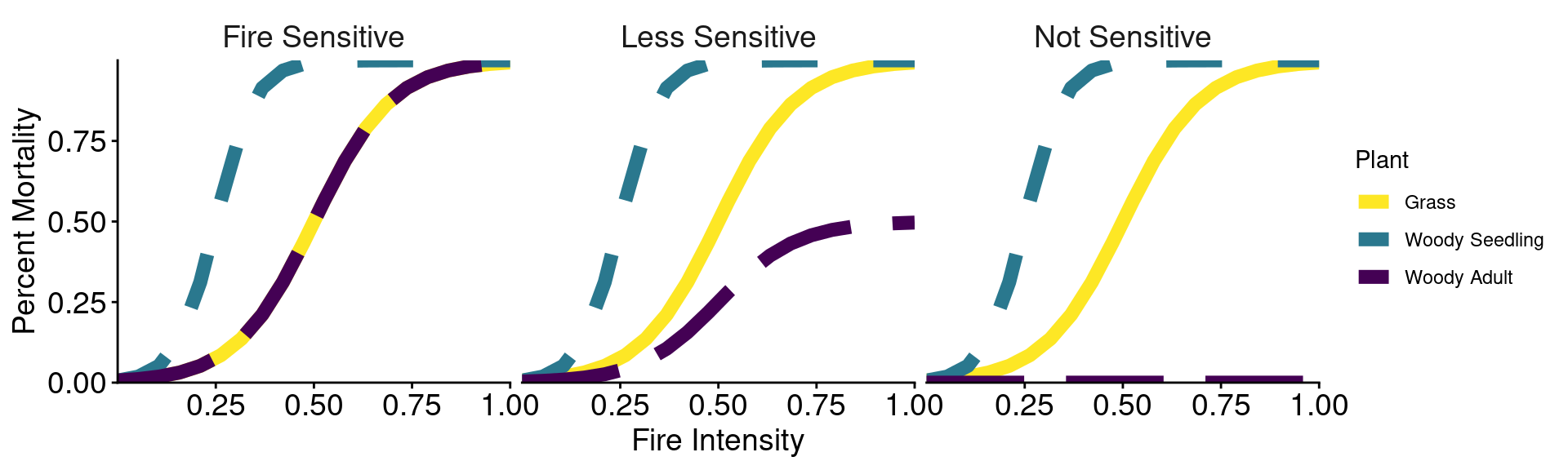
Model predictions with fire
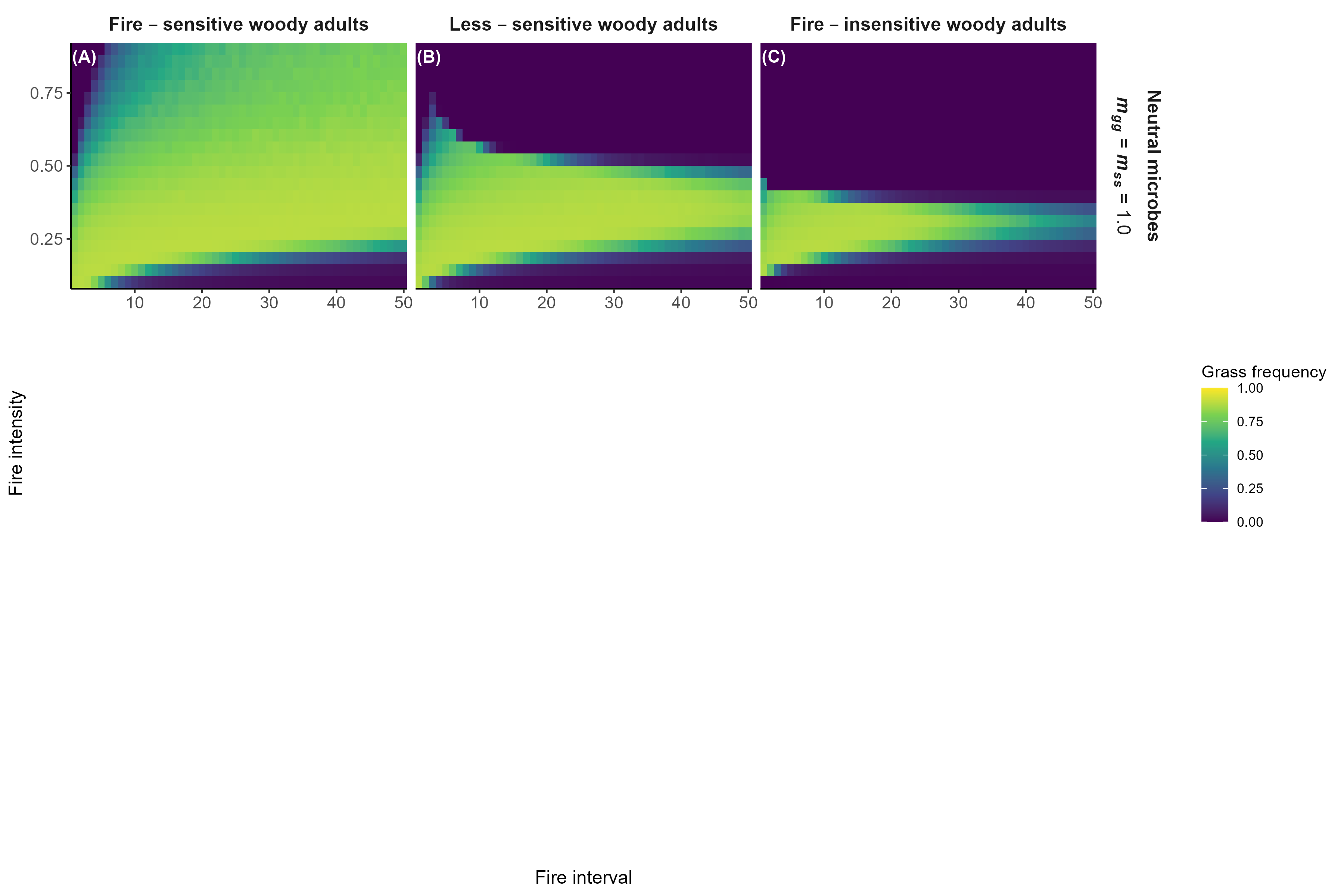
Model predictions with fire
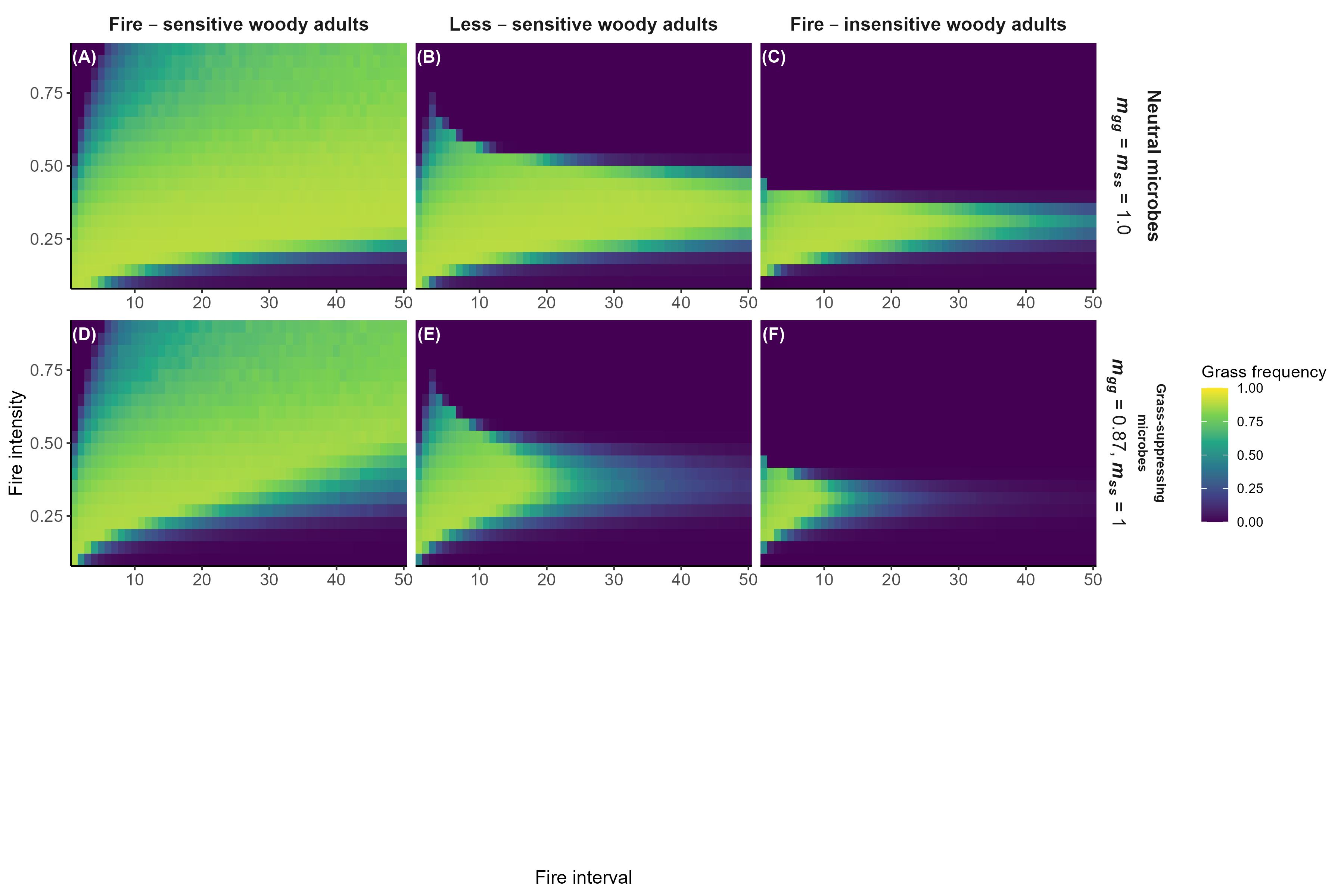
Model predictions with fire
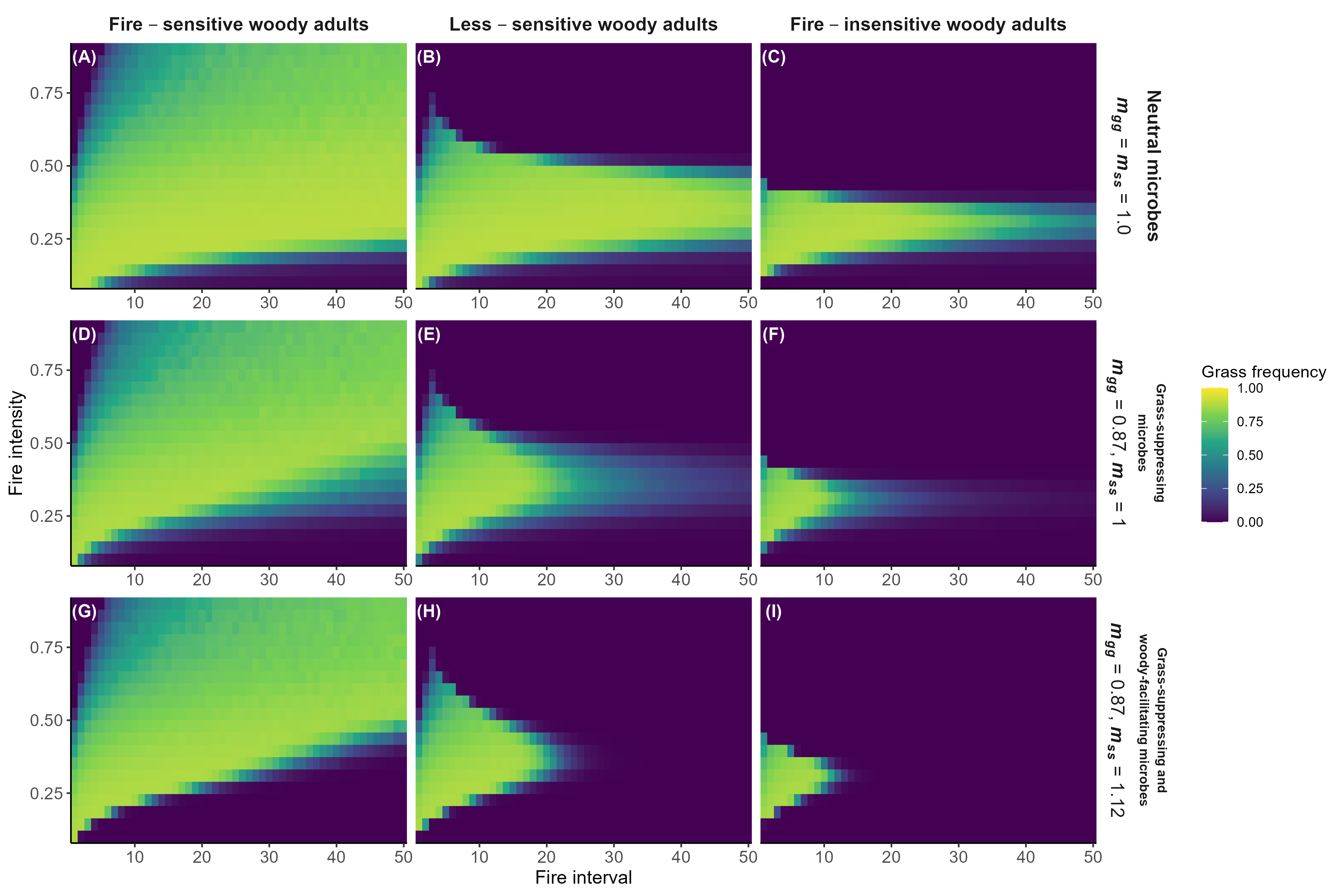
Interpretation:
Microbial feedbacks constrain which fire regimes sustain grassy communities
Microbial feedbacks can also constrain fire-based grassland restoration after woody encroachment
Detailed in preprint (Simha and Kandlikar 2026), feedback welcome
Historical perspective on fire in longleaf pine

Interrogating the human legacies of fires (and fire suppression)
Working in collaboration with the Southeastern Tribal Alliance for Fire (SETA Fire), we hope to…
- Document variation in cultural burn practices
- Explore stories about fire, soil, and biodiversity
- Contrast the practice and impacts of cultural vs. prescribed burns
Today’s talk
Ecological theory for microbial effects on plant coexistence
Microbial effects on plant communities in variable environments
Other ongoing projects:
- Coexistence and species invasions in fragmented tropical forests (Western Ghats of India)
- Microbial contributions to plant eco-evolutionary responses to drought
Today’s talk
Ecological theory for microbial effects on plant coexistence
Microbial effects on plant communities in variable environments
- Reflections on broadening representation and equity
- Outlook under the second Trump administration
Survey of scientist sentiments
Developed a survey to ask about life scientists’ sentiments of the second Trump administration’s policies, particularly:
- Impacts of immigration policy;
- Impacts of funding policy;
- Impacts on science as a whole.
160 responses, mostly from tenured faculty who are US citizens.
Thematic analysis of open-ended responses
Note: We ran the survey from May-September 2025.
Impacts of immigration policy
Although practically all responses were from US citizens, many expressed deep concerns for self, colleagues, and mentees.
“I definitely feel more racially-profiled than I used to before this administration.”
Immigration policies are also changing professional behaviors:
- Many plan to stop taking on international mentees (some even say it would be unethical to do so)
Impacts of funding policy
Most responses focused on grant cancellations and the threat of future cancellations
Some respondents articulated their strategies: Alter language in research proposals, pursue new directions where possible.
Others report anticipating a complete devastation of their research programs.
Financial strain also manifests in personal uncertainty.
PIs report profound stress that accompanies the responsibility for lab members’ livelihoods
Uncertain implications for career prospects, even for those with tenure-track positions
Impacts on science as a whole
Concern for training pipeline, including international scholars (“brain drain”)
Concerns of anti-science sentiment and politicization of science
Fears of the decline of the US as a global science leader
Pervasive feelings of helplessness and that the losses are irrevocable
How to transform individual helplessness into strategic and collective action?
- Some food for thought, reflecting on the survey responses
Nelson (2026) offers a useful framework:
“When resources contract, the most responsible response is […] to ask what we are actually trying to protect. Is it every individual project, or is it the capacity of our research communities to keep asking meaningful questions, train thinkers and preserve what we already know?”
How do these priorities shape data collection, training, hiring, evaluation?
History reveals that what we are up against is a culmination of long-term trends.
History reveals that what we are up against is a culmination of long-term trends.
“The choice of research directions is made on the basis of marketability […]. Support is most likely for projects that promise a secure outcome that fits the agendas of its sponsors.”
“[T]the allocation of resources within a university is influenced by the prestige and earning capacity of the various programs, and scientists report pressure [to] turn their research in more affluent directions, such as genetic engineering.”
Levins and Lewontin, “Biology under the influence” (2007) and “Dialectical biologist” (1987)
What science do we want on the “other side”?
- The systems of science have been under strain for a long time.
Yet there have been signs of movement:
- Scientists as a group are more diverse than ever before.
- Groups like BREWS exist that aim to take science in this new direction.

Thank you!
slides: https://talks.gklab.org/umn-26
contact: gkandlikar@lsu.edu