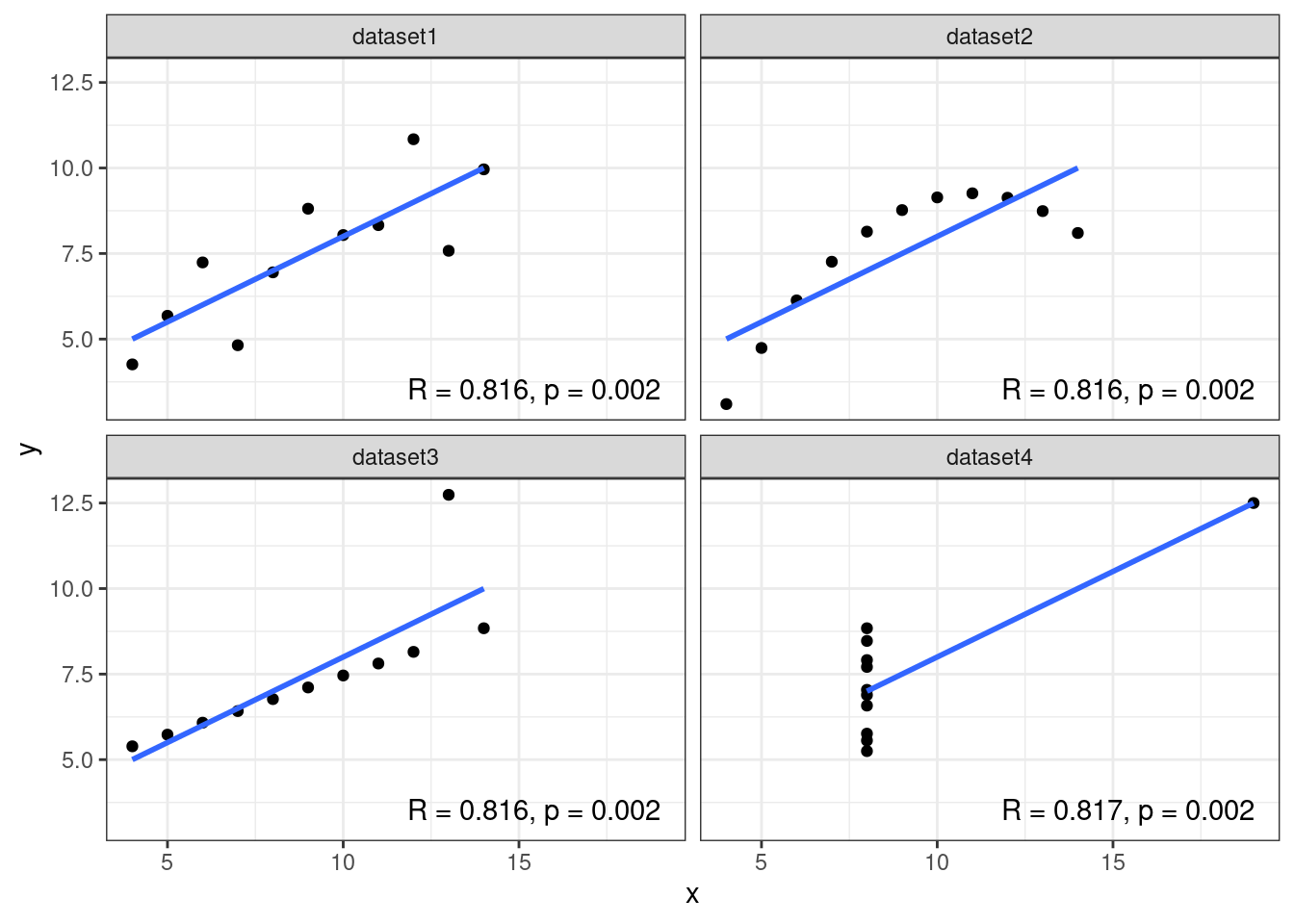
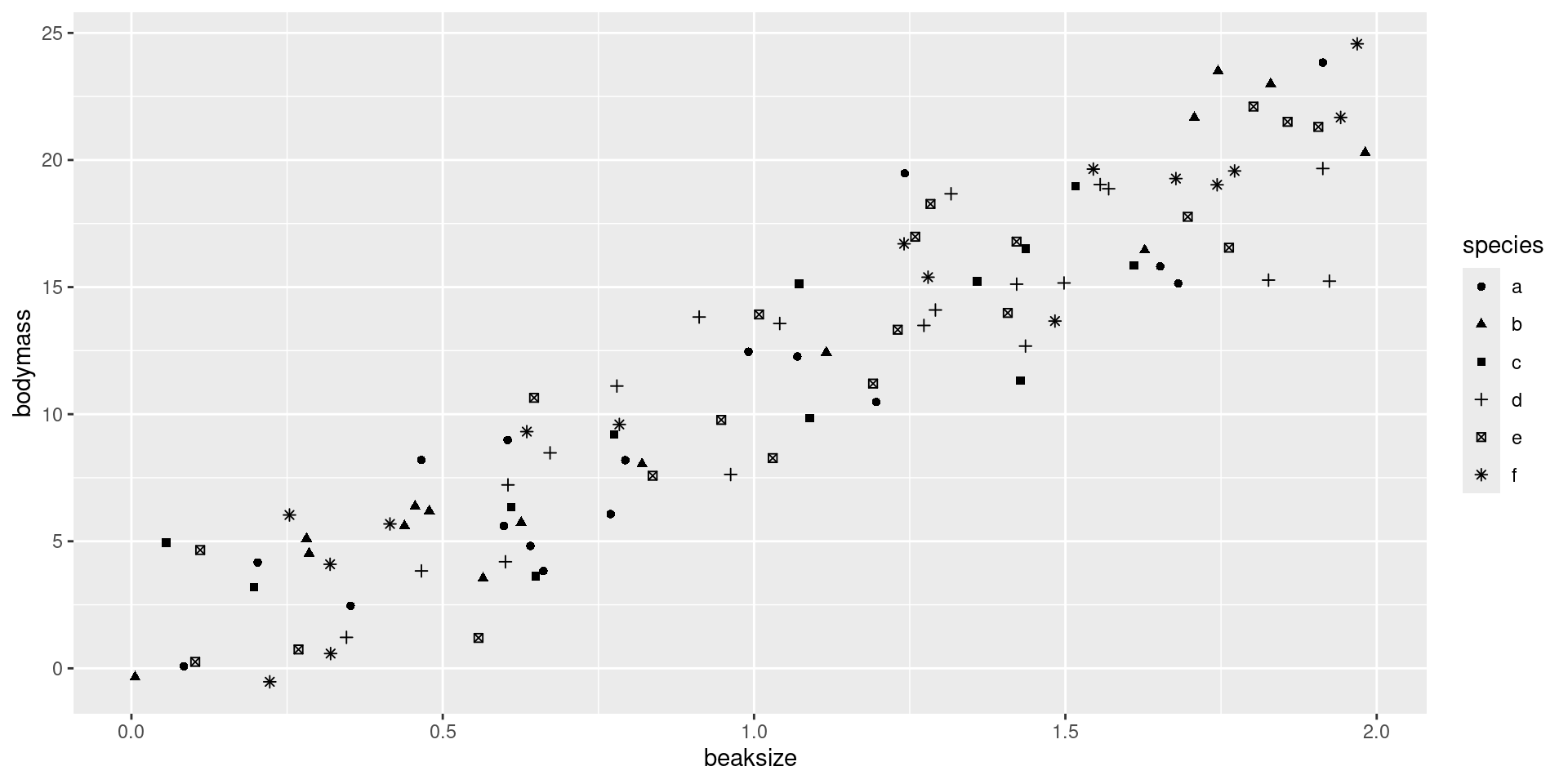
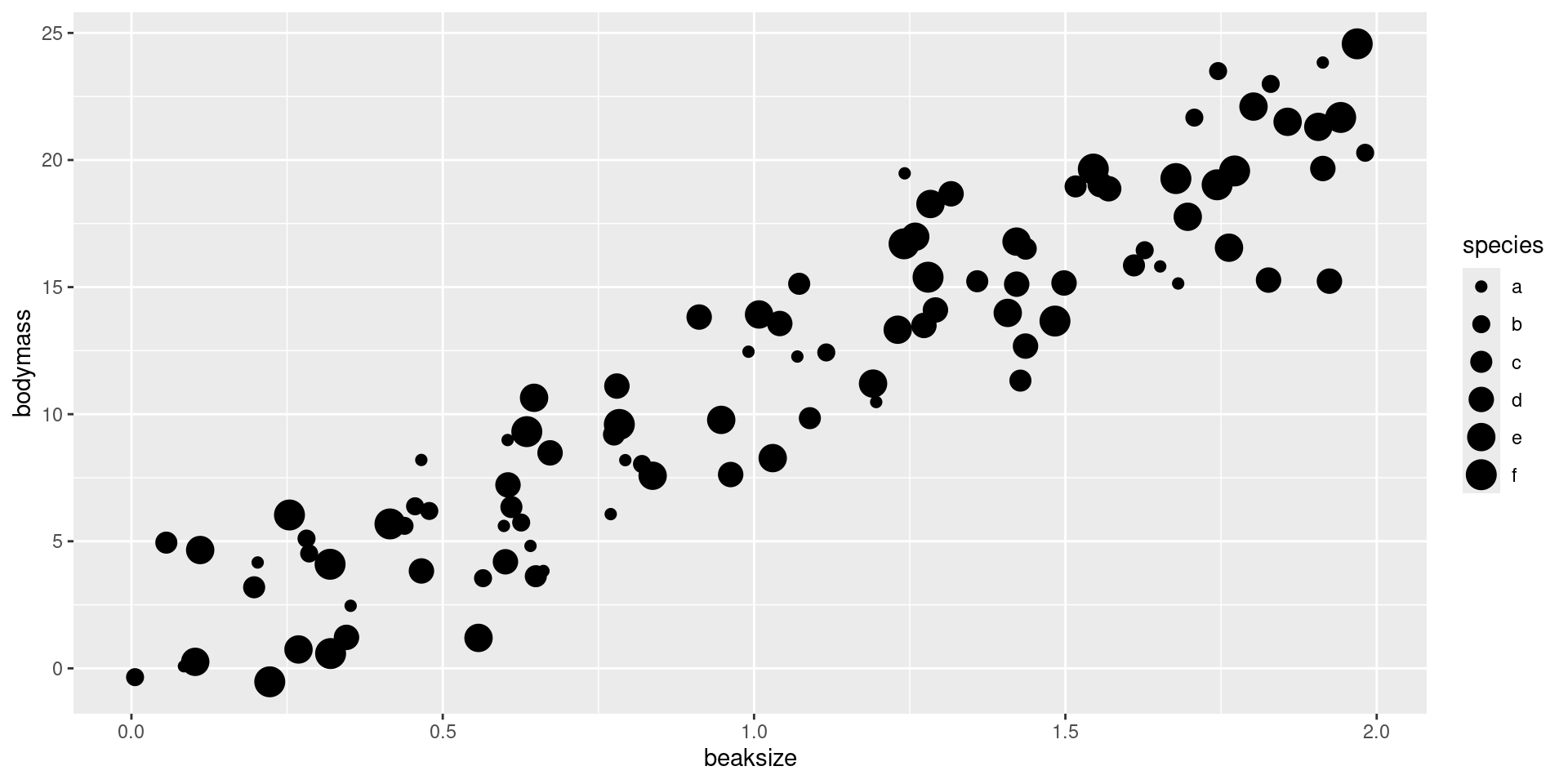
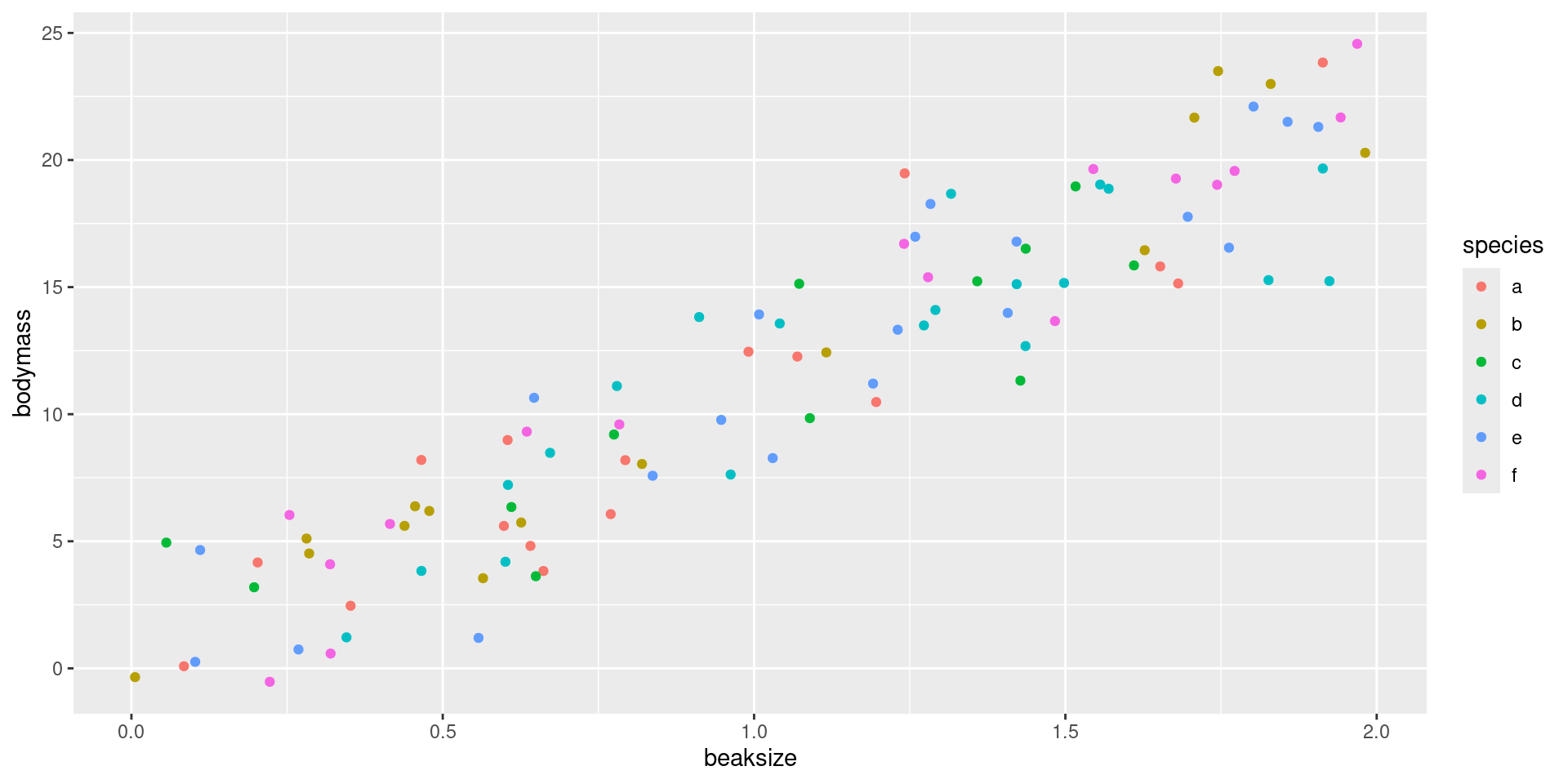
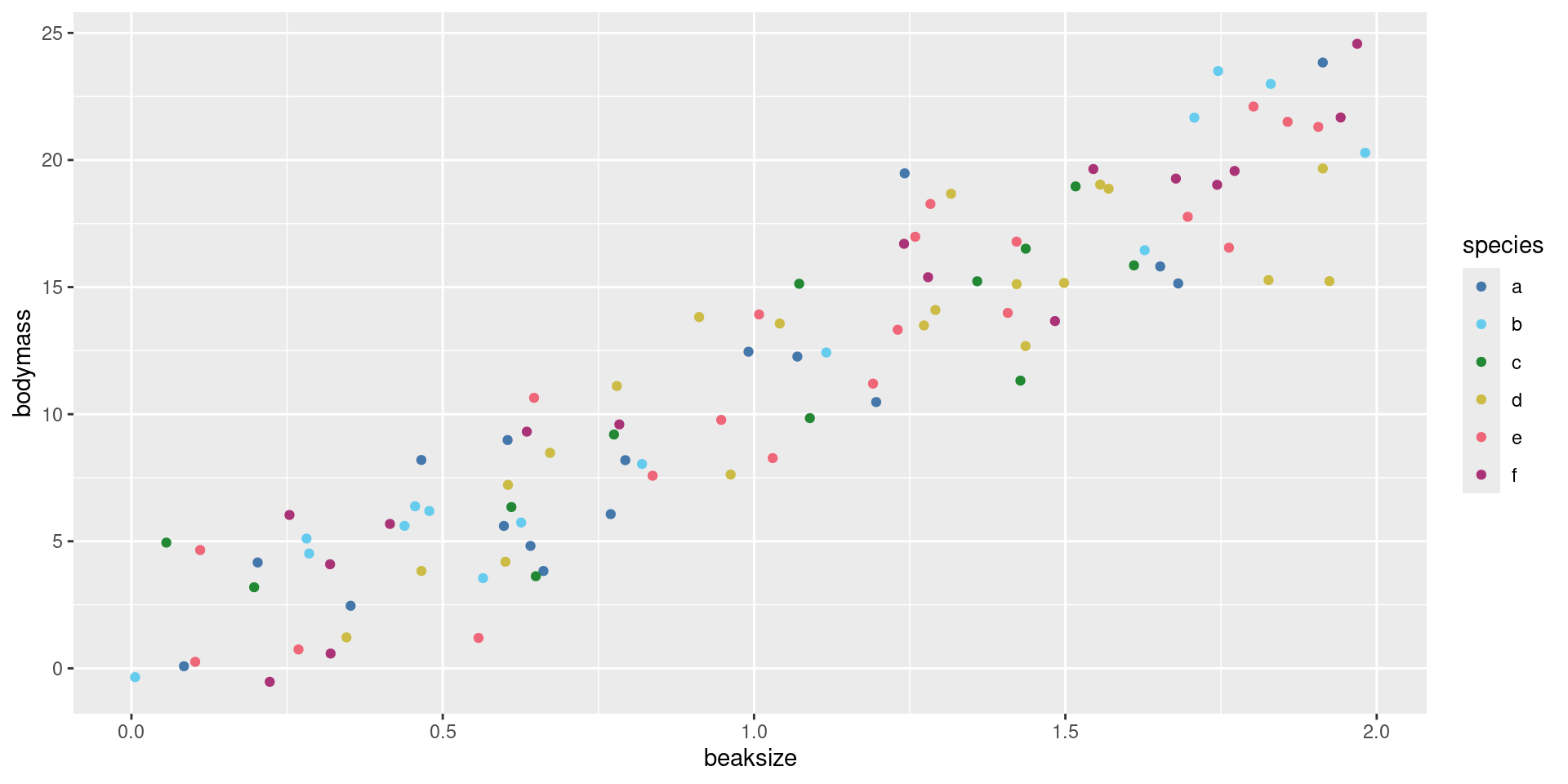
After assigning each country-year to one of four public debt/GDP groups, RR calculates the average real GDP growth for each country within the group, that is, a single average value for the country for all the years it appeared in the category.
For example, real GDP growth in the UK averaged 2.4 percent per year during the 19 years that the UK appeared in the highest public debt/GDP category while real GDP growth for the US averaged −2.0 percent per year during the 4 years that the US appeared in the highest category.
Both were weighed equally in the final analysis, despite one country contributing 19 years of data, vs. 4 years for the other.